Institute of Oceanology, Chinese Academy of Sciences
Article Information
- LI Yingdong(李应东), XUE Huiling(薛慧玲), LI Xiaodong(李晓东)
- Transcriptome analysis of the Chinese grass shrimp Palaemonetes sinensis (Sollaud 1911) and its predicted feeding habit
- Chinese Journal of Oceanology and Limnology, 36(5): 1778-1787
- http://dx.doi.org/10.1007/s00343-019-7189-y
Article History
- Received Jun. 30, 2017
- accepted in principle Sep. 20, 2017
- accepted for publication Nov. 13, 2017
The Chinese grass shrimp Palaemonetes sinensis is a small shrimp (2–5 cm long) belonging to the Palaemonidae family of decapod crustaceans and is widely distributed in China and adjacent areas (Li et al., 2007). Many of them are non-native and have been introduced locally (Bradbury et al., 1996; Cai and Dai, 1999; Imai and Oonuki, 2014). Palaemonetes sinensis is an important aquarium animal and fisheries resource because of its striking appearance and appealing flavor (unpublished data). It is also commonly sold as bait for sport fishing (Imai and Oonuki, 2014). Palaemonetes sinensis has been historically consumed by humans in China and has a strong market value, and often reaches prices higher than those for penaeid species. However, resources of P. sinensis are rapidly decreasing because of habitat destruction and overfishing. To date, biological knowledge of this species is scarce, and studies have mainly focused on their morphological analysis (Imai and Oonuki, 2014). Given the importance of studies in a wide variety of contexts, sufficient genomic tools would greatly enhance all aspects of research on this species, especially those on functional gene discovery, molecular marker application, and genome comparison. To our knowledge, neither publicly available sequence data nor functional genetic information of P. sinensis have yet been reported.
Transcriptome sequencing has been shown to be an effective means of gene discovery, especially when no genome sequence is available (Koboldt et al., 2013). We sequenced the first P. sinensis transcriptome from the whole body of several individuals. Additionally, the study of digestive enzymes enables better comprehension of the mechanism of digestion and the nutritional needs to be applied in the formulation of inert diets, which are essential for profitable aquaculture; therefore, we aimed to identify the genes related to digestive enzymes. Finally, we developed a set of polymorphic microsatellites and estimated the genetic diversity of a wild P. sinensis population. The resulting transcriptome will be useful in gene function demonstration, molecular markerassisted selection, and other genetic studies on P. sinensis and related shrimp species.
2 MATERIAL AND METHOD 2.1 Ethics statementOur study does not involve endangered or protected species. In China, catching wild grass shrimp from rice fields does not require specific permits. Experimental treatment of the Chinese grass shrimp in this study was conducted according to national and international guidelines. The experimental protocol was approved by the Animal Ethics Committee of Shenyang Agriculture University.
2.2 Sampling of experimental animalsA total of 10 adult P. sinensis shrimps were collected randomly from a rice field in Panjin City, Liaoning Province, China, in October 2015. Body size of the shrimps ranged from 2.56 to 3.61 cm, with a mean of 3.04±0.34 cm SD. Individual shrimps were placed in a 2-mL RNAse-free tube and immediately placed into liquid nitrogen prior to RNA isolation.
2.3 RNA preparation and sequencingTotal RNA was extracted from each of the 10 individual shrimps using the Trizol Kit (Invitrogen, USA) following the manufacturer`s instructions. RNA quality and quantity were examined using 1% TAE agarose gel electrophoresis. Equal quantities (1 μg) of RNA obtained from each shrimp were pooled together to eliminate variation among the samples and used as one sample for RNA-Seq library construction. The library evaluated in this study was assembled using NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA) following the protocol of the manufacturer. Library fragments were purified using the AMPure XP system (Beckman Coulter, Beverly, USA) in order to select cDNA fragments with lengths of 150–200 bp. Three microliters of USER Enzyme (NEB, USA) were added to a 180–480-bp pooled sample of size-selected, adaptor-ligated cDNA at 37℃ for 15 min, followed by 5 min at 95℃. Next, PCR was performed with Phusion High-Fidelity DNA polymerase with GC buffer and 2 mol/L betaine, Universal PCR primers and Index (X) Primer. Finally, PCR products were purified using the AMPure XP system and the library quality was assessed on the Agilent Bioanalyzer 2100 system. The mixed DNA libraries were diluted to 4–5 pmol/L for sequencing by the Illumina Hiseq 2500TM platform.
2.4 De novo assembly and gene annotationClean reads were obtained by filtering out lowquality sequences and adapter sequences from the raw reads. To obtain complete reference sequences, clean reads were assembled using the Trinity program. Contigs with longer fragments were further assembled form clean reads until they could not be extended at either side. Unigenes were obtained by removing the assembled redundant transcripts.
All assembled transcripts were aligned with the NCBI protein databases NR (non-redundant), PFAM (protein family), Swiss-Prot (a manually annotated and reviewed protein sequence database), KOG/COG (Clusters of Orthologous Groups of proteins), KEGG (Kyoto Encyclopedia of Genes and Genomes), KO (KEGG ortholog database) and GO (Gene Ontology) using BLASTx (with a cutoff E-value of 1E-05). The best BLAST hits from these databases were then selected for further functional annotation of the unigenes. Gene ontology (GO) annotation was analyzed by Blast2GO program in order to obtain GO annotations of unique assembled transcripts.
2.5 SSR DiscoveryUnique sequences were scanned for potential simple sequence repeat (SSR) markers using the MISA software (http://pgrc.ipk-gatersleben.de/misa) with default parameters. SSRs were searched for with motifs of sizes ranging from mono- to hexanucleotides, with repeat units of SSRs of 6, 5, 4, 3 and 2, respectively. Primer design was performed using the Batch Primer 3 software.
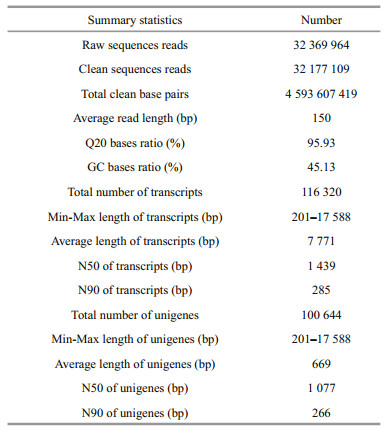
3 RESULT 3.1 Transcriptome sequencing and de novo assemblyAll raw reads have been submitted to the GenBank database under accession number SRR1630824. A total of 32 369 964 raw reads and 32 177 109 clean reads were obtained from whole P. sinensis. A total of 100 644 unigenes are generated with N50 length of 1 077 bp and average length of 667 bp (Table 1). The size distribution of the shrimp transcripts and unigenes were presented in Supplementary file (Fig.S1).
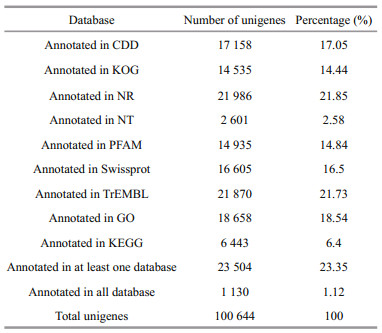
All unigenes were compared to eight public databases using BLASTx with an E-value threshold of 10-5. Approximately 23 504 (23.35% of 100 644) unigenes showed significant BLAST hits with one or more of the databases (Table 2). For most of the reads in the NR database, similar unigenes were retrieved from Daphnia pulex (1 985, 9.03%), Tribolium castaneum (1 068, 4.86%), Vittaforma corneae (800, 3.64%), Pediculus humanus corporis (762, 3.47%), Prunus persica (723, 3.29%), Strongylocentrotus purpuratus (643, 2.92%), Branchiostoma floridae (579, 2.63%), Ixodes scapularis (518, 2.36%), Nasonia vitripennis (496, 2.26%), and other species (10 302, 46.86%) (Fig.S2).
Three categories were defined by the GO project, including biological processes, molecular function, and cellular components. A total of 18 658 unigenes were assigned to GO categories (Fig. 1), with 17 298 unigenes assigned to more than one category. The top five most frequent categories of biological processes in our study were "cellular processes" (13.13%), "metabolic processes" (10.93%), "single-organism processes" (8.43%), "biological regulation" (6.1%), and "regulation of biological processes" (5.66%).

|
| Figure 1 Gene Ontology (GO) terms for the transcriptomic sequences of Palaemonetes sinensis |
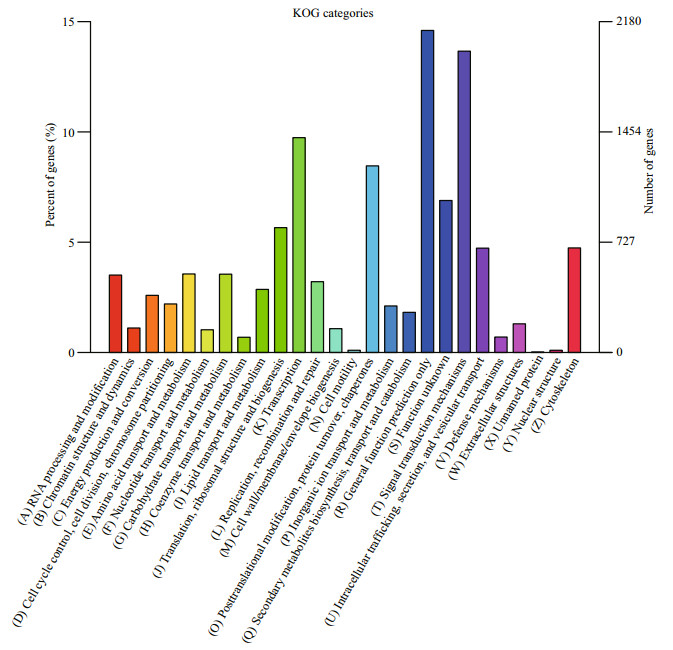
To predict and classify their possible functions with reference to orthologs from other species, all unigenes were annotated by the Eukaryotic Orthologous Groups (KOG) database. As shown in Fig. 2, a total of 14 536 (61.84%) unigenes were categorized into 25 groups. "General function prediction only" accounted for the largest group (2 122, 14.60%), followed by "signal transduction mechanisms" (1 986, 13.66%) and "transcription" (1 415, 9.73%). "Carbohydrate transport and metabolism" accounted for 516 unigenes and 518 unigenes were assigned to "amino acid transport and metabolism."
|
| Figure 2 KOG classification of putative proteins for Palaemonetes sinensis transcriptome |
To further elucidate the biological functions based on gene interactions, all unigenes were mapped against the KEGG pathway database. A total of 6 443 unigenes were assigned to 338 pathways (Table S1). The most enriched pathways were "metabolism" (2 705, 41.98%), "organismal systems" (2 043, 31.71%), "genetic information processing" (1 676, 26.01%), "environmental information processing" (1 187, 18.42%), and "cellular processing" (1 053, 16.34%) (Fig. 3).

|
| Figure 3 Function distribution of all unigenes in the transcriptome of Palaemonetes sinensis based on KEGG analysis |
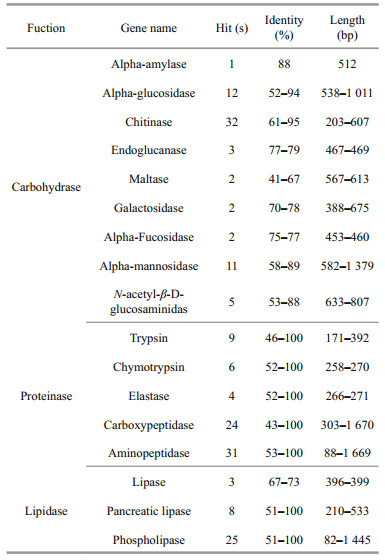
In this study, 180 unigenes were classified as being involved with the Chinese grass shrimp digestive enzyme system: carbohydrase, proteinase, and lipidase (Table 3). Nine categories of carbohydrase were identified, which were composed of highly expressed alpha-glucosidase, alpha-mannosidase, and chitinase enzyme genes with over 10 hits. Thirty-two chitinase unigenes were identified in P. sinensis, the highest number for any carbohydrase (45.71%). A total of 19 serine proteinase genes were identified, including nine trypsin, six chymotrypsin and four elastase unigenes. In addition, we also identified 24 carboxypeptidase and 31 aminopeptidase unigenes (Table S2). A total of 36 lipidase-related unigenes were identified in this study, consisting of three lipase unigenes, eight pancreatic lipase unigenes, and 25 phospholipase unigenes.
|
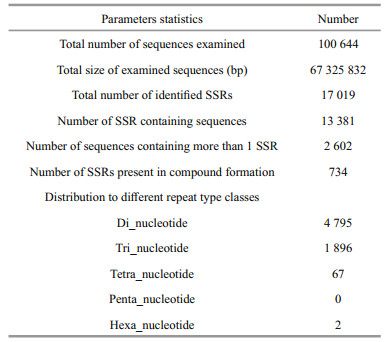
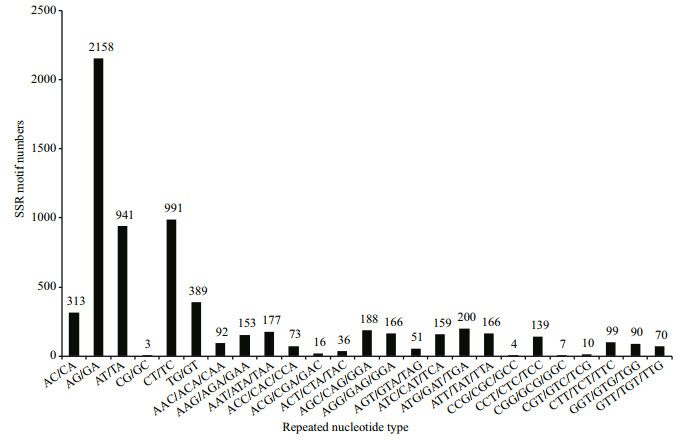
We obtained a total of 17 019 SSRs in the transcriptomic dataset, followed by 28.17% dinucleotide, 11.14% tri-nucleotide and 0.41% tetra/ penta/hexa-nucleotide repeats (Table 4). The most common types among the 20 types of tri-nucleotide repeats were (ATG/GAT/TGA) n, (AGC/CAG/GCA) n, and (AAT/ATA/TAA) n (Fig. 4).
|
| Figure 4 Distribution of simple sequence repeat (SSR) nucleotide classes among different nucleotide types found in Palaemonetes sinensis |
Information on the genomic resources of crustaceans is still appreciably lacking, unlike that on insects and vertebrates. Transcriptome sequencing has been widely used and provides a viable alternative to characterizing complete genomes (Havird and Santos, 2016). After Griffitt et al. (2007) provided the first serial analysis of gene expression (SAGE) in grass shrimp Palaemonetes gugio, García et al. (2017) provided the first de novo transcriptome data of grass shrimp Palaemonetes argentines and identified 24 378 transcripts. In our current study, we have provided a total of 116 320 transcripts of P. sinensis shrimp transcriptome data using Illumina Hiseq 2500TM platform. The average length of all transcripts and N50 were 771 bp and 1 439, respectively, which were longer than those in P. argentines (504 bp and 621). Long sequences of good quality enabled us to obtain more information about genes. Moreover, we also obtained a total of 17 019 SSRs in the transcriptomic dataset. The SSR number of P. sinensis was smaller than that in most crabs such as Portunus trituberculatus (22 673), Scylla paramamosain (19 011) and Eriocheir sinensis (141 737), but larger than that in shrimps such as Macrobrachium nipponense (6 689) and Fenneropenaeus chinensis (14 981). This information will provide a basis for future studies on the molecular biology of P. sinensis.
Although the activity and composition of digestive enzymes have been assessed in many crustacean species (Figueiredo and Anderson, 2009; Ayalaborboa et al., 2013; Sacristán et al., 2015), the corresponding molecular information is still lacking. Therefore, identification of genes related to digestive enzymes is useful and helps to establish the functioning of the crustacean digestive system in relation to the nutritional requirements and feeding habits at the molecular level. Unigenes related to digestive enzymes were highly represented in this study. A total of 180 unigenes were identified, which were composed of 69 carbohydrases, 74 proteinases, and 36 lipidases (Table 3). However, Wang et al. (2014) and Dammannagoda et al. (2015) identified only 114 and 139 digestive enzyme genes from the hepatopancreatic transcripts of P. trituberculatus and Cherax quadricarinatus, respectively. Although the hepatopancreas is the main organ for digestive enzyme synthesis in the crustacean digestive system, other digestive organs, such as the intestine and stomach can also synthesize digestive enzymes (Shi et al., 2009; Proespraiwong et al., 2010; Ayalaborboa et al., 2013). Therefore, transcriptomes of whole digestive systems are needed to identify all the digestive enzyme genes. Because it was difficult to separate the digestive apparatus manually from such small a shrimp (only 2–5 cm), the whole body of P. sinensis was used in this study.
Carbohydrase is crucial for carbohydrate metabolism in crustaceans (Perera et al., 2008; Jiang et al., 2009), and plays an important role in triggering the utilization of protein and lipid for energy production (Holme et al., 2009; Gucic et al., 2013). According to our sequence analysis, nine categories of carbohydrase were identified, of which alphaglucosidase, alpha-mannosidase, and chitinase enzyme genes were expressed highly with over 10 hits (Table 3). Chitinolytic enzymes are involved in digestion of chitin-containing food, and they are present in many crustaceans (Kono et al., 1995). In our study, 32 chitinase unigenes were identified in P. sinensis, the highest number for any carbohydrases (45.71%), which showed close similarity with the reported chitinase sequences in the morotoge shrimp Pandalopsis japonica (Table S2).
Commercial shrimp feeds are protein-rich in comparison with other aquatic animal feeds (Martinez-Cordova et al., 2003; Cavalli et al., 2004). However, there is scarcity of information on the protein requirements of Palaemonidae species (Palma et al., 2015). Given their relatively high protein requirements, the initial focus of protein requirement studies should be proteinases. In most studies, the presence of these enzymes has only been detected in raw extracts, using synthetic substrates and specific inhibitors for each enzyme. In the present study, a total of 19 serine proteinase genes were identified, including nine trypsin, six chymotrypsin, and four elastase unigenes. In addition, we also identified 24 carboxypeptidase and 31 aminopeptidase unigenes (Table S2). Dietary lipids play an important role in crustacean nutrition, by providing essential fatty acids and energy. However, lipid digestion has not been studied in the Palaemonidae species. A total of 36 lipidase-related unigenes were identified in this study, including three lipase unigenes, eight pancreatic lipase unigenes, and 25 phospholipase unigenes.
A better understanding of the crustacean feeding habits will contribute to increasing the performance of crustacean breeding programs (Varadharajan and Pushparajan, 2012). In general, Palaemonetes are opportunistic omnivores, feeding on the littoral benthic communities (Collins, 1999; Aguzzi et al., 2005). However, a previous study has shown that P. sinensis prefers to eat meiofauna and dead tissues from conspecifics (unpublished data). It is understood that the feeding habits of crustaceans have close relationship with the structure and function of their digestive enzymes (Aragón-Axomulco et al., 2012). Therefore, herbivorous species like Macrobrachium australiense show high levels of carbohydrase and low levels of protease. Carnivorous species like Portunus pelagicus and Scylla serrata show an opposite trend in the levels of these enzymes in their bodies (Figueiredo and Anderson, 2009). Our transcriptome data from P. sinensis indicates that the number of carbohydrase unigenes (69 hits) is similar to the number of proteinase unigenes (74 hits). It appears that the feeding habits of P. sinensis reflects an omnivorous or detritivorous habit. Additionally, Broad (1957) found that the larvae of both Palaemonetes pugio and Palaemonetes vulgaris were unable to survive if fed on unicellular algae alone. Moreover, the feeding strategy of Palaemonetes antennarius in the laboratory involves facultative specialization on dead and live animals rather than plant detritus (Costantini and Rossi, 2001).
Alpha-amylase and endo-β-mannanase, the main carbohydrases in crustaceans, allow crustaceans to digest food from plant sources. In our study, only one genomic sequence of alpha-amylase and none of endo-β-mannanase were identified; these values are lower than that reported from herbivorous or omnivorous decapod species (Van Wormhoudt et al., 1995; Dammannagoda et al., 2015; Rodríguez-Viera et al., 2016). These results indicate that P. sinensis may be a carnivorous or saprophagous species.
Endoglucanase is a key enzyme for the degradation of cellulose and allows animals to digest diverse foods from plant sources (Smant et al., 1998). The endoglucanase genes have been identified in mussels (Xu et al., 2001), clams (Nikapitiya et al., 2010), and red crayfish (Byrne et al., 1999). Dammannagoda et al. (2015) identified 48 endoglucanase genes from the hepatopancreatic transcripts of Cherax quadricarinatus. In our study, only three endoglucanase unigenes were identified to be related to cellulose hydrolysis. C. quadricarinatus more efficiently consume and digest vegetal ingredients than animal ingredients (Campaña-Torres et al., 2005; Torres et al., 2006), explaining the greater number of its endoglucanase genes. In addition, many studies suggested that there may be at least some endogenous cytolytic enzymes in saprophagous invertebrates (Treves and Martin, 1994; Watanabe et al., 1997; Smant et al., 1998). The molecular discovery of endoglucanase genes in this study indicates that P. sinensis has the ability to digest cellulose, supporting the hypothesis of a saprophagous habit.
Trypsin enzymes play a central role in protein digestion in crustaceans. In C. quadricarinatus, six trypsin-related unigenes were identified, which is not much lesser than that in P. sinensis. In P. trituberculatus, 44 trypsin-related unigenes were identified. The number of proteinase unigenes identified from P. sinensis were the same as that identified from C. quadricarinatus (Dammannagoda et al., 2015). Nonetheless, further research is needed to consolidate the present knowledge on protein requirements of this shrimp.
5 CONCLUSIONThe current study presents the first comprehensive high-throughput dataset obtained for the Chinese grass shrimp P. sinensis. A total of 100 644 unigenes and 17 019 simple sequence repeats were identified. Moreover, 180 unigenes were classified as being involved with the Chinese grass shrimp digestive enzyme system, including proteinase, carbohydrase, lipidase, as well as other related important genes. The first transcriptomic investigation of P. sinensis may provide abundant genomic data and offer comprehensive sequence resources for further elaborate studies on the grass shrimp. Additionally, the data generated here will contribute to the study of digestive enzymes, and may help in understanding the links between digestive enzymes and food habit. Hence, further research on digestive enzyme genes is essential, with focus on important aspects like the levels and patterns of expression of endoglucanase, alpha-amylase and endo-β-mannanase.
6 DATA AVAILABILITY STATEMENTAll sequence data that support the findings of this study have been deposited in the NCBI Short Read Archive (SRA) with the primary accession code https://www.ncbi.nlm.nih.gov/sra/SRX1956097.
7 ACKNOWLEDGEMENTWe thank Panjin Guanghe fisheries Co., Ltd. for providing the shrimps.
Electronic supplementary materialSupplementary material (Supplementary Tables S1–S2, Fig.S1–S2) is available in the online version of this article at https://doi.org/10.1007/s00343-019-7189-y.
Aguzzi J, Cuesta J A, Librero M, Toja J. 2005. Daily and seasonal feeding rhythmicity of Palaemonetes varians (Leach 1814) from southwestern Europe. Marine Biology, 148(1): 141-147.
DOI:10.1007/s00227-005-0025-2 |
Aragón-Axomulco H, Chiappa-Carrara X, Soto L, Cuzon G, Arena L, Maldonado C, Cárdenas R, Gaxiola G. 2012. Seasonal variability in trypsin and α-amylase activities caused by the molting cycle and feeding habits of juvenile pink shrimp Farfantepenaeus duorarum (Burkenroad, 1939). Journal of Crustacean Biology, 32(1): 89-99.
DOI:10.1163/193724011X615361 |
Ayalaborboa E G, Tovarramírez D, Civeracerecedo R, Rojascontreras M, Cadenaroa M A, Nolascosoria H, Goytortúabores E. 2013. Effect of red crab meal (Pleuroncodes planipes) on growth and digestive enzyme expression in the intestine of white shrimp (Litopenaeus vannamel). Communications in Agricultural and Applied Biological Sciences, 78(4): 19-22.
|
Bradbury P C, Zhang L M, Shi X B. 1996. A redescription of Gymnodinioides caridinae (Miyashita 1933) from Palaemonetes sinensis (Sollaud 1911) in the Songhua River. The Journal of Eukaryotic Microbiology, 43(5): 404-408.
DOI:10.1111/jeu.1996.43.issue-5 |
Broad A C. 1957. The relationship between diet and larval development of palaemonetes. Biological Bulletin, 112(2): 162-170.
DOI:10.2307/1539194 |
Byrne K A, Lehnert S A, Johnson S E, Moore S S. 1999. Isolation of a cDNA encoding a putative cellulase in the red claw crayfish Cherax quadricarinatus. Gene, 239(2): 317-324.
DOI:10.1016/S0378-1119(99)00396-0 |
Cai Y, Dai A Y. 1999. Freshwater shrimps (Crustacea: Decapoda:Caridea) from the Xishuangbanna region of Yunnan Province, southern China. Hydrobiologia, 400: 211-241.
DOI:10.1023/A:1003717109973 |
Campaña-Torres A, Martinez-Cordova L R, Villarreal-Colmenares H, Civera-Cerecedo R. 2005. In vivo dry matter and protein digestibility of three plant-derived and four animal-derived feedstuffs and diets for juvenile Australian redclaw, Cherax quadricarinatus. Aquaculture, 250(3-4): 748-754.
DOI:10.1016/j.aquaculture.2005.02.058 |
Cavalli R O, Zimmermann S, Speck R C. 2004. Growth and feed utilization of the shrimp Farfantepenaeus paulensis fed diets containing different marine protein sources. Ciência Rural, 34(3): 891-896.
DOI:10.1590/S0103-84782004000300036 |
Collins P A. 1999. Feeding of Palaemonetes argentinus (Decapoda:Palaemonidae) from an oxbow lake of the paraná river, Argentina. Journal of Crustacean Biology, 19(3): 485-492.
DOI:10.2307/1549257 |
Costantini M L, Rossi L. 2001. Laboratory study of the grass shrimp feeding preferences. Hydrobiologia, 443(1-3): 129-136.
|
Dammannagoda L K, Pavasovic A, Prentis P J, Hurwood D A, Mather P B. 2015. Expression and characterization of digestive enzyme genes from hepatopancreatic transcripts from redclaw crayfish (Cherax quadricarinatus). Aquaculture Nutrition, 21(6): 868-880.
DOI:10.1111/anu.12211 |
Figueiredo M S R B, Anderson A J. 2009. Digestive enzyme spectra in crustacean decapods (Paleomonidae, Portunidae and Penaeidae) feeding in the natural habitat. Aquaculture Research, 40(3): 282-291.
DOI:10.1111/are.2009.40.issue-3 |
García C F, Pedrini N, Sánchez-Paz A, Reyna-Blanco C S, Lavarias S, Muhlia-Almazán A, Fernández-Giménez A, Laino A, De-La-Re-Vega E, Lukaszewicz G, LópezZavala A A, Brieba L G, Criscitello M F, CarrascoMiranda J S, García-Orozco K D, Ochoa-Leyva A, Rudiño-Piñera E, Sanchez-Flores A, Sotelo-Mundo R R. 2017. De novo assembly and transcriptome characterization of the freshwater prawn Palaemonetes argentinus: implications for a detoxification response. Marine Genomics, https://doi.org10.1016/j.margen.2017.08.009.
|
Griffitt R J, Greig T W, Chandler G T, Quattro J M. 2007. Serial analysis of gene expression reveals identifiable patterns in transcriptome profiles of Palaemonetes pugio exposed to three common environmental stressors. Environmental Toxicology and Chemistry, 26(11): 2 413-2 419.
DOI:10.1897/07-158R.1 |
Gucic M, Cortés-Jacinto E, Civera-Cerecedo R, Ricque-Marie D, Martínez-Córdova L R. 2013. Apparent carbohydrate and lipid digestibility of feeds for whiteleg shrimp, Litopenaeus vannamei (Decapoda:Penaeidae), cultivated at different salinities. Revista de Biologia Tropical, 61(3): 1 201-1 213.
|
Havird J C, Santos S R. 2016. Here we are, but where do we go? a systematic review of crustacean transcriptomic studies from 2014-2015. Integrative and Comparative Biology, 56(6): 1 055-1 066.
DOI:10.1093/icb/icw061 |
Holme M H, Zeng C S, Southgate P C. 2009. A review of recent progress toward development of a formulated microbound diet for mud crab, Scylla serrata, larvae and their nutritional requirements. Aquaculture, 286(3-4): 164-175.
DOI:10.1016/j.aquaculture.2008.09.021 |
Imai T, Oonuki T. 2014. Records of Chinese grass shrimp, Palaemonetes sinensis (Sollaud, 1911) from western Japan and simple differentiation method with native freshwater shrimp, Palaemon paucidens De Haan, 1844 using eye size and carapace color pattern. Bioinvasions Record, 3(3): 163-168.
DOI:10.3391/bir |
Jiang H, Yin Y X, Zhang X W, Hu S N, Wang Q. 2009. Chasing relationships between nutrition and reproduction:a comparative transcriptome analysis of hepatopancreas and testis from Eriocheir sinensis. Comparative Biochemistry and Physiology Part D:Genomics and Proteomics, 4(3): 227-234.
DOI:10.1016/j.cbd.2009.05.001 |
Koboldt D C, Steinberg K M, Larson D E, Wilson R K, Mardis E R. 2013. The next-generation sequencing revolution and its impact on genomics. Cell, 155(1): 27-38.
DOI:10.1016/j.cell.2013.09.006 |
Kono M, Wilder M N, Matsui T, Furukawa K, Koga D, Aida K. 1995. Chitinolytic enzyme activities in the hepatopancreas, tail fan and hemolymph of kuruma prawn Penaeus japonicus during the molt cycle. Fisheries Science, 61(4): 727-728.
DOI:10.2331/fishsci.61.727 |
Li X Z, Liu R Y, Liang X Q, Chen G. 2007. Fauna finica. Invertebrate Vol. 44, Subphylum Crustacea, Order Decopoda, Superfamily Palaemonoidea. Science Press, Beijing, China.
(in Chinese)
|
Martinez-Cordova L R, Torres A C, Porchas-Cornejo M A. 2003. Dietary protein level and natural food management in the culture of blue (Litopenaeus stylirostris) and white shrimp (Litopenaeus vannamei) in microcosms. Aquaculture Nutrition, 9(3): 155-160.
DOI:10.1046/j.1365-2095.2003.00235.x |
Nikapitiya C, Oh C, De Zoysa M, Whang I, Kang D H, Lee S R, Kim S J, Lee J. 2010. Characterization of beta-1, 4-endoglucanase as a polysaccharide-degrading digestive enzyme from disk abalone, Haliotis discus discus. Aquaculture International, 18(6): 1 061-1 078.
DOI:10.1007/s10499-010-9324-9 |
Palma J, Andrade J P, Lemme A, Bureau D P. 2015. Quantitative dietary requirement of juvenile Atlantic ditch shrimp Palaemonetes varians for lysine, methionine and arginine. Aquaculture Research, 46(8): 1 822-1 830.
DOI:10.1111/are.2015.46.issue-8 |
Perera E, Moyano F J, Díaz M, Perdomo-Morales R, MonteroAlejo V, Alonso E, Carrillo O, Galich G S. 2008. Polymorphism and partial characterization of digestive enzymes in the spiny lobster Panulirus argus. Comparative Biochemistry and Physiology Part B:Biochemistry and Molecular Biology, 150(3): 247-254.
DOI:10.1016/j.cbpb.2008.03.009 |
Rodríguez-Viera L, Perera E, Martos-Sitcha J A, PerdomoMorales R, Casuso A, Montero-Alejo V, García-Galano T, Martínez-Rodríguez G, Mancera J M. 2016. Molecular, biochemical, and dietary regulation features of α-amylase in a carnivorous crustacean, the spiny lobster Panulirus argus. PLoS One, 11(7): e0158919.
DOI:10.1371/journal.pone.0158919 |
Proespraiwong P, Tassanakajon A, Rimphanitchayakit V. 2010. Chitinases from the black tiger shrimp Penaeus monodon: Phylogenetics, expression and activities. Comparative Biochemistry and Physiology Part B:Biochemistry and Molecular Biology, 156(2): 86-96.
DOI:10.1016/j.cbpb.2010.02.007 |
Sacristán H J, Fernández-Gimenez A V, Chaulet A, Tadic L M F, Fenucci J, Greco L S L. 2015. Effect of different diets on digestive enzyme activities, in vitro digestibility, and midgut gland structure in juvenile crayfish, Cherax quadricarinatus. Acta Zoologica, 97(4): 407-418.
|
Shi X Z, Ren Q, Zhao X F, Wang J X. 2009. Expression of four trypsin-like serine proteases from the Chinese shrimp, Fenneropenaeus chinensis, as regulated by pathogenic infection. Comparative Biochemistry and Physiology Part B:Biochemistry and Molecular Biology, 153(1): 54-60.
DOI:10.1016/j.cbpb.2009.01.011 |
Smant G, Stokkermans J P W G, Yan Y T, De Boer J M, Baum T J, Wang X H, Hussey R S, Gommers F J, Henrissat B, Davis E L, Helder J, Schots A, Bakker J. 1998. Endogenous cellulases in animals:isolation of β-1, 4-endoglucanase genes from two species of plant-parasitic cyst nematodes. Proceedings of the National Academy of Sciences of the United States of America, 95(9): 4 906-4 911.
DOI:10.1073/pnas.95.9.4906 |
Torres A C, Córdova L R M, Colmenares H V, Cerecedo R C. 2006. Apparent dry matter and protein digestibility of vegetal and animal ingredients and diets for pre-adult Australian redclaw crayfish Cherax quadricarinatus (Von Martens 1858). Hidrobioló gica, 16(1): 103-106.
|
Treves D S, Martin M M. 1994. Cellulose digestion in primitive hexapods:effect of ingested antibiotics on gut microbial populations and gut cellulase levels in the firebrat, Thermobia domestica (Zygentoma, Lepismatidae). Journal of Chemical Ecology, 20(8): 2 003-2 020.
DOI:10.1007/BF02066239 |
Van Wormhoudt A, Bourreau G, Le Moullac G. 1995. Amylase polymorphism in crustacea decapoda:electrophoretic and immunological studies. Biochemical Systematics and Ecology, 23(2): 139-149.
DOI:10.1016/0305-1978(94)00090-4 |
Varadharajan D, Pushparajan N. 2012. Food and feeding habits of aquaculture candidate a potential Crustacean of Pacific white shrimp Litopenaeus vannamei, South East Coast of India. Journal of Aquaculture Research & Development, 4: 161.
|
Wang W, Wu X G, Liu Z J, Zheng H J, Cheng Y X. 2014. Insights into hepatopancreatic functions for nutrition metabolism and ovarian development in the crab Portunus trituberculatus:gene discovery in the comparative transcriptome of different hepatopancreas stages. PLoS One, 9(1): e84921.
DOI:10.1371/journal.pone.0084921 |
Watanabe H, Nakamura M, Tokuda G, Yamaoka I, Scrivener A M, Noda H. 1997. Site of secretion and properties of endogenous endo-β-1, 4-glucanase components from Reticulitermes speratus (Kolbe), a Japanese subterranean termite. Insect Biochemistry and Molecular Biology, 27(4): 305-313.
DOI:10.1016/S0965-1748(97)00003-9 |
Xu B Z, Janson J C, Sellos D. 2001. Cloning and sequencing of a molluscan endo-β-1, 4-glucanase gene from the blue mussel, Mytilus edulis. European Journal of Biochemistry, 268(13): 3 718-3 727.
DOI:10.1046/j.1432-1327.2001.02280.x |
 2018, Vol. 36
2018, Vol. 36