Institute of Oceanology, Chinese Academy of Sciences
Article Information
- WANG Qi(王琦), SHI Yu(石禹), HE Maoxian(何毛贤)
- Pf-Dmrt4, a potential factor in sexual development in the pearl oyster Pinctada fucata
- Chinese Journal of Oceanology and Limnology, 36(6): 2337-2350
- http://dx.doi.org/10.1007/s00343-019-7177-2
Article History
- Received Jun. 19, 2017
- accepted in principle Aug. 20, 2017
- accepted for publication Dec. 27, 2017
2 Taiyuan University of Technology, Taiyuan 030024, China
Sexual reproduction is a remarkable phenomenon in biology and the primary source of genetic variation required for evolutionary survival. It can be found in almost all types of multicellular animals. The mechanisms of sex determination and differentiation vary among phyla, possibly as a result of long-term evolution of the genes involved in the sex determination pathways (Marı́n and Baker, 1998). Bivalves belong to the second largest phylum and display highly diverse sexual phenotypes. The mollusc pearl oyster Pinctada fucata exhibits complex modes of sexual reproduction that consists of protandric dioecy, occasional sex change and hermaphroditism. Environmental factors such as population density, water temperature, and sex ratios play important roles in sex determination and differentiation in bivalves (Yu et al., 2007, 2011), though the function of genetic factors remains unclear.
The Drosophila doublesex (Burtis and Baker, 1989) and Caenorhabditis elegans mab-3 (Shen and Hodgkin, 1988) genes are widely conserved and have been implicated in sex determination (Raymond et al., 1999). The Dmrt (double-sex and mab-3-related transcription factor) gene family encodes proteins with conserved DM domains (a common zinc fingerlike DNA-binding motif) and is represented by multiple members in both vertebrates and invertebrates (Volff et al., 2003). To date, eight members of the family have been reported in humans (Ottolenghi et al., 2002) and seven have been found in mice (Kim et al., 2003). DM-domain family members have been shown to regulate sexual development in vertebrates and invertebrates, and the function of the Dmrt gene family in sex regulation has been highly conserved during evolution (Hodgkin, 2002).
Apart from Dmrt1, DM-domain family members are expressed during development in tissues other than the gonads, suggesting that these proteins may control a broader range of developmental processes (Hong et al., 2007). Some Dmrt4 and Dmrt5 orthologs play key roles in sexual development in different animals such as zebrafish (Guo et al., 2004; Winkler et al., 2004), blue tilapia (Cao et al., 2010), Chlamys farreri (Feng et al., 2010) and Crassostrea. gigas (Naimi et al., 2009).
As a highly conserved gene family, Dmrt genes have been cloned and studied in various vertebrates (Ottolenghi et al., 2002; Aoyama et al., 2003) and several invertebrates, including fruit flies (Crémazy et al., 2001), nematodes (Volff et al., 2003), sea squirts (Leveugle et al., 2004) and freshwater prawns (Peng et al., 2005). In molluscs, orthologs of the DMdomain transcription factors have been isolated from the oyster C. gigas (CgDMl) (Naimi et al., 2009), the scallop Ch. farreri (Cfdmrt4-like) (Feng et al., 2010), the abalone Haliotis asinina (HADMRT1-like) (Klinbunga et al., 2009), the pearl oyster Pinctada martensii (pmDmrt2 and pmDmrt5) (Yu et al., 2009, 2011), and the scallop Ch. nobilis (dmrt5) (Shi et al., 2014). Gene expression analysis of some of these orthologs suggests the possible involvement of these genes in gonad differentiation and development (Naimi et al., 2009; Feng et al., 2010). Sex determination and differentiation in molluscs appear to be complex because of the diverse range of species and underlying molecular mechanisms involved.
Recently, the genomics of some economically marine bivalves are studied to provide better understanding of the molecular mechanisms underlying their different reproductive strategies such as the black-lip pearl oyster P. margaritifera (Teaniniuraitemoana et al., 2014), the Pacific oyster C. gigas (Zhang et al., 2014) and the pearl oyster P. fucata (Takeuchi et al., 2012; Du et al., 2017). The available transcriptome is a powerful and novel resource to study sexual pathway in marine mollusks.
It is an essential step to identify and characterize sex-specific genes to explore the molecular pathways of sexual development in the pearl oyster P. fucata. This study therefore investigated the genetic mechanisms responsible for sex differentiation and gonad development of Pf-Dmrt4 in P. fucata, which is an economically important species for pearl production in China. The full-length cDNA sequence of Pf-Dmrt4 during gonad differentiation was characterized, and the spatio-temporal expression patters of Pf-Dmrt4 were studied during gonad differentiation and in adult tissue using real-time quantitative reverse transcription-polymerase chain reaction (qRT-PCR). The cellular localization and the sex-steroid -induced expression of the gene were also investigated.
2 MATERIAL AND METHOD 2.1 Experimental animals and tissue samplingPearl oysters P. fucata were obtained from Daya Bay in Shenzhen (Guangdong Province, China), cultivated in floating net cages in the sea under natural conditions. All specimens were collected between May, 2012 and August, 2013. Testes and ovaries were excised, frozen immediately in liquid nitrogen and maintained at -80℃ until RNA extraction for Pf-Dmrt4 cloning. Ovary, testis, heart, digestive gland, adductor muscle, mantle, and gill tissues were also dissected and stored in the same manner for tissuedistribution analysis. Gonad tissues from different stages were dissected, fixed in Bouin's solution at 4℃ overnight, and submitted to histological observation to determine the gonadal stages. Gonad samples were fixed with 4% paraformaldehyde dissolved in phosphate-buffered saline (PBS) overnight at 4℃ and then stored in methanol at -20℃ for in- situ hybridization. Gametes were obtained and passed through a 100-μm screen to remove tissue debris. The eggs were mixed with sperms at 25-26℃ in filtered seawater which contained 0.005%-0.006% (v/v) ammonia. After fertilization, embryos and different stages of larvae were then prepared for RNA extraction.
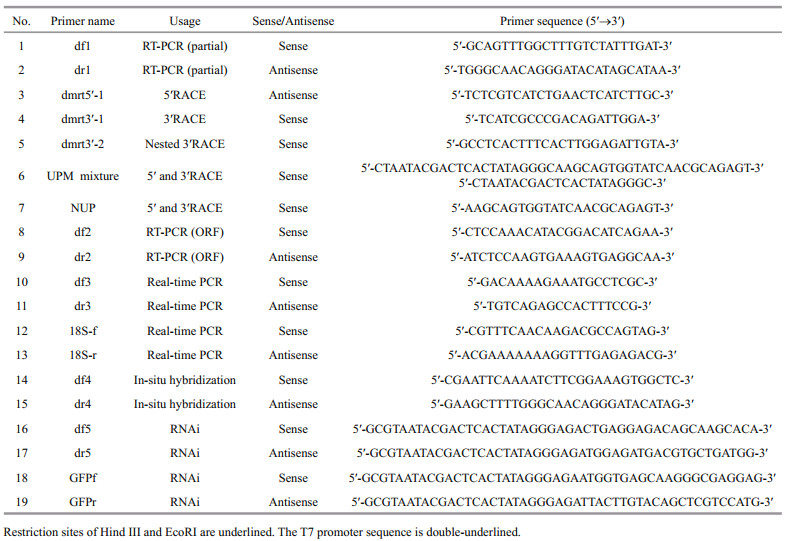
2.2 Cloning and sequencing of Pf-Dmrt4Total RNA was extracted from testes using an E.Z.N.A. Mollusc RNA Kit according to the manufacturer's protocol (Omega, Japan). The quantity and quality of RNA were assayed by UV spectrophotometry (OD260/OD280). A ReverTra AceFirst-strand cDNA Synthesis Kit (Toyobo, Japan) was used to synthesize first-strand cDNA from 1 μg total RNA. All primers for Pf-Dmrt4 used in the present study are listed in Table 1. PCR primers were designed using Primer Premier 5.00 (Premier Biosoft International, Palo Alto, CA, USA). All primer pairs were designed to eliminate false-positive bands of potential genomic DNA contamination. A fragment obtained from the transcriptome of P. fucata was amplified using a pair of specific primers df 1 and dr1. BD SMART RACE cDNA Amplification Kit (Clontech, USA) was used to synthesize SMART cDNA. The resulting 1 384-bp fragment was used to generate full-length cDNA by 5′- and 3′ rapid amplification of cDNA ends. The cycling conditions for all PCR were: 3 min at 94℃ for denaturation, 35-40 cycles at 94℃ for 30 s, 55-60℃ for 30 s and 72℃ for 1-2 min, followed by a further elongation at 72℃ for 10 min. The PCR products were purified with The E.Z.N.A. Gel Extraction Kit (Omega BioTek, USA), subcloned into the pGEM-T Easy Vector (Promega, Madison, WI, USA) and sequenced (Applied Biosystems, USA).
Tissue expression patterns of Pf-Dmrt4 mRNA were analyzed by real-time qPCR. Expression analysis was performed in ovary, testis, gill, mantle, heart, digestive gland, and adductor muscle tissues collected from three pearl oysters and stored at -80℃. Reactions were performed in triplicate. Real-time qPCR were carried out using a Roche LightCycler 480 real-time PCR system, with SYBR green master mix (Toyobo, Japan) following the manufacturer's instructions. All the qPCR primers are presented in Table 1. Real-time PCR was performed as follows: 94℃ for 1 min, and 40 cycles of 94℃ for 15 s, 15 s at 55℃ and 72℃ for 1 min.
2.4 Expression patterns of Pf-Dmrt4 in embryonic and larval P. fucataThe developmental expression patterns of Pf-Dmrt4 were analyzed in five developmental stages, which were collected and stored at -80℃: blastulastage embryo, trochophore, D-shaped larva, umbo larva and metamorphosis stage. Total RNA extraction and real-time qPCR were the same as described above and three repetitions were analyzed.
2.5 Expression patterns of Pf-Dmrt4 in P. fucata gonads during gametogenesisGonadal sections were fixed with Bouin's fixative, 6 μm slices were prepared, and processed with conventional histological analysis. The sections were stained with hematoxylin-eosin and examined microscopically to classify the natural gonadal development. The gametogenic expression patterns of Pf-Dmrt4 were analysed by real-time PCR in cDNAs from four stages of ovary and testis (growing stage, mature stage, spawning stage and resting stage), synthesized as described above. Five replicates of each sample were analyzed.
2.6 Expression of Pf-Dmrt4 mRNA following methyltestosterone injectionThe effect of methyltestosterone (MT) on the mRNA expression levels of Pf-Dmrt4 were investigated by injecting male oysters with MT (purchased from Guangzhou Lifescience Biotechnology Co. Ltd., Guangzhou, China) at the gonadal mature stage. MT was dissolved in dimethyl sulfoxide (DMSO) (Sigma, St. Louis, MO) and added into modified Herbst's artificial seawater (ASW) to a concentration of 1 μg/μL. A 100-μL aliquot of suspension was injected into the adductor muscle of male oysters. The animals in Negative control group were injected with 100 μL of ASW containing 1% DMSO. Eight oysters were randomly collected before injection (blank control). A further eight oysters were collected every 6 h after injection for 3 times, respectively, and the gonads were dissected and fixed for RNA extraction and real-time PCR.
2.7 In situ hybridizationThe expression of Pf-Dmrt4 in gonads were localized using the gene-specific primers designed for amplification (Table 1). Digoxigenin -labeled RNA probes were prepared from a 482-bp fragment of the Pf-Dmrt4 cDNA that had been amplified and subcloned into pGEM-T Easy vectors (Promega) as a template. The probes were labeled with a DIG RNA labelling kit (Roche Diagnostics, Switzerland). Transcriptions were performed using SP6 (sense) or T7 (antisense) RNA polymerase. Paraformaldehydefixed gonad samples were embedded in paraffin, sectioned (10 μm), de-paraffinized, and then rehydrated. In situ hybridization was performed as described by Miyashita et al. (2008), with some modifications. Gonad sections were hybridized with DIG-labeled RNA probes, and hybridized signals were visualized with NBT/BCIP system and alkaline phosphatase-conjugated anti-DIG antibody using a DIG Nucleic Acid Detection Kit (Roche Diagnostics) following the manufacturer's protocol.
2.8 RNA interferenceRNA interference was performed as described by Suzuki et al. (2009), with some modifications. The primers used to generate Pf-Dmrt4 and green fluorescent protein (GFP) double-stranded (ds) RNAs are shown in Table 1. The T7 promoter sequence is underlined. RiboMAXTM Large Scale RNA Production System (T7) kit (Promega, USA) was uesd to synthesize and purify the dsRNA. RNase-free DNase I (TaKaRa, Japan) was used to digest template DNA. Synthesized dsRNA was diluted to 10, 20 and 40 μg/100 μL with PBS, and 100 μL of each solution was injected into the adductor muscles of 2-year-old oysters with mature testes. PBS 100 μL or 20 μg GFP dsRNA in 100 μL PBS were used as blank and negative controls. Each treatment selected 7 oysters. Total RNA was extracted from the gonad of each individual 7 days after injection and used for firststrand cDNA synthesis. The expression levels were quantified by real-time qPCR, using 18S as a reference gene. The Pf-Dmrt4 qPCR primers were designed to amplify the region that did not overlap the region targeted by dsRNA. Gonads were dissected from the injected-oyster groups 7 days after injection, fixed in Bouin's solution at 4 ℃ overnight, and submitted to histological observation at the light-microscope level.
2.9 Statistical analysisHomology analysis of the deduced amino acid (aa) sequence was conducted using the public NCBIBLAST database. Motif analysis was performed using InterPro Scan on the ExPASy Proteomics Server. ClustalX (1.81) was used for multiple sequence alignments of amio acids. A phylogenetic tree was constructed by the neighbor-joining method using MEGA.6. Three-dimensional model of PfDMRT4 were predicted by SWISS Model on http://swissmodel.expasy.org/. All reference sequences were downloaded from GenBank and public genome resources. Quantitative data were shown as mean±S.E.M. Significant differences were estimated by one-way ANOVA followed by Duncan's multiple range tests.
3 RESULT 3.1 Isolation and characterization of Pf-Dmrt4A full-length cDNA encoding Pf-Dmrt4 was isolated from P. fucata (GenBank accession number: KM272582). The cDNA sequence of Pf-Dmrt4 was 1 711 bp, containing an open reading frame of 1 104 bp, 5′ untranlated region (UTR) of 199 bp and 3′UTR of 408 bp with a 30-bp polyA tail. The deduced aa sequence was 367 aa and contained a DM domain consensus sequence (from 23-81 aa) with conserved cysteines and histidines characteristic of the DMRT protein family, and a DMA domain (from 207-242 aa) (Fig. 1).

|
| Fig.1 Nucleotide and deduced amino acid sequences of Pf-Dmrt4 DM domain is boxed, the underlined and the dashed lines indicate the DMA domain and the short motif, respectively. The cysteine and histidine conserved residues are bolded and underlined. |
The amino acid sequence of the conserved DM domain of P. fucata Pf-DMRT4 was aligned with known sequences in various members of this family in different species (Fig. 2a). The highest identity rate was 100% with P. martensii DMRT5, followed by Ch. farreri DMRT4-like, C. gigas DMI and Ch. nobilis DMRT5 (98%). Sequence comparison revealed a conserved zinc module consisting of intertwined CCHC and HCCC Zn2+-binding sites, and a consensus sequence of five aa (KGHKR) of the putative nuclear localization signal (NLS). There was a conserved DMA domain and a short conserved domain of seven aa (KSAFSPI) near the C-terminus in DMRT4 and DMRT5, but not in DMRT-like types (Fig. 2b). The DMA domain of P. fucata DMRT4 displayed highest homologies of 74% with Cg-DMI, 68% with Ch. farreri DMRT4-like, 66% with Ch. nobilis DMRT5 and 65% with Oryzias latipes DMRT5. The seven-aa conserved domain shared the highest levels of aa identity with Ch. farreri DMRT4-like and Ch. nobilis DMRT5 (100%), Cg-DMI and H. sapiens DMRT4 (85.71%) and H. sapiens DMRT5 (71.43%).

|
| Fig.2 Alignment of amino acid sequences of the DM domain of P. fucata DMRT4 with other DMRTs of different species (a) and alignment of amino acid sequences of the DMA domain and the short conserved motif from different species (b) In a: identical residues are shown in black, dark and light gray represent strongly or weakly conserved similar amino acids. The short conserved signal NLS (Nuclear Localization Signal) (KGHKR) is underlined and the zinc module consisting of intertwined CCHC and HCCC Zn2+-binding sites (Site Ⅰ and Ⅱ) are indicated with black boxes. In b: identical amino acids to Pf-Dmrt4 is underlined. The numbers of the amino acid residues between DMA domain and the short conserved motif are indicated in the black box. (GenBank accession numbers were listed in Supplementary Material 1). |
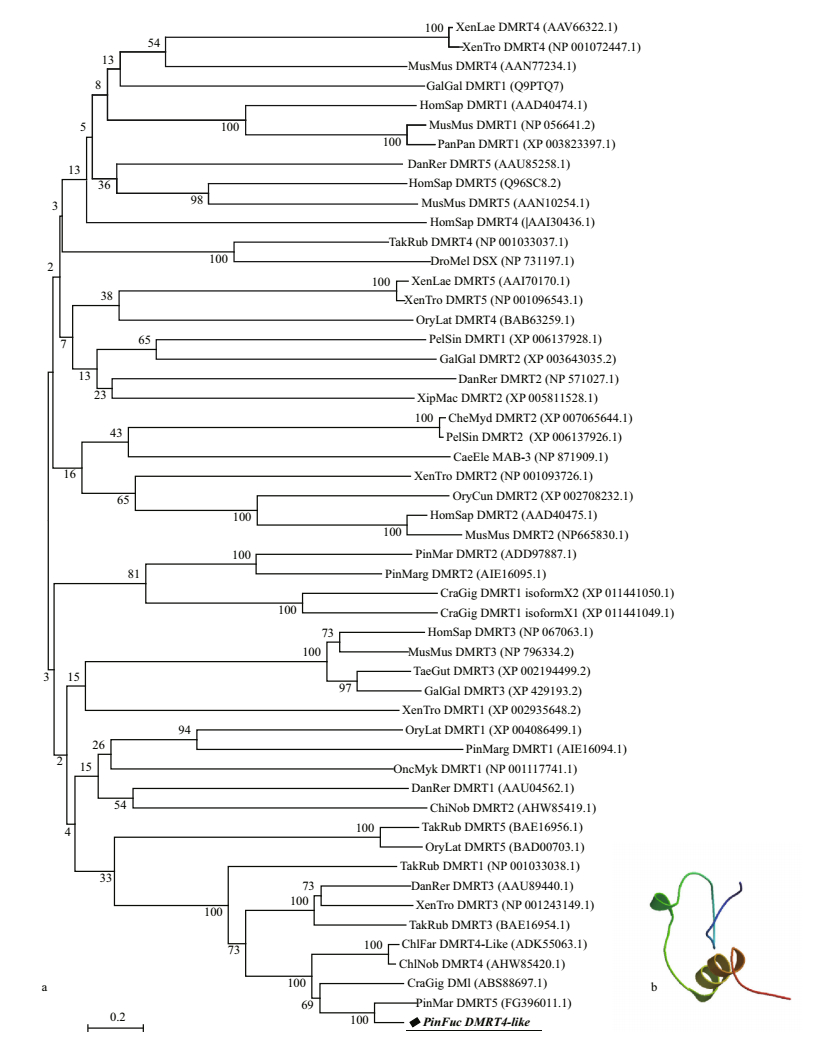
The phylogenetic tree generated using the complete protein genes of DMRT from various species showed that P. fucata DMRT4 was most closely related to P. martensii DMRT5, then clustered with Ch. farreri DMRT4-like, C. gigas DMI and Ch. nobilis DMRT5 (Fig. 3a). P. fucata DMRT4 was grouped with the mollusc DMRT4 and DMRT5 clusters, but more closely with DMRT4, implying that P. fucata DMRT4 may be a member of the DMRT4 cluster. Two transcripts encoding orthologs of the DM domain transcription factor were identified in P. margaritifera, PinMarg DMRT1 and PinMarg DMRT2 were clustered with OryLat DMRT1 and PinMar DMRT2, then grouped with DMRT1 isoform X1 and X2 which were found in the genomic database in C. gigas.
|
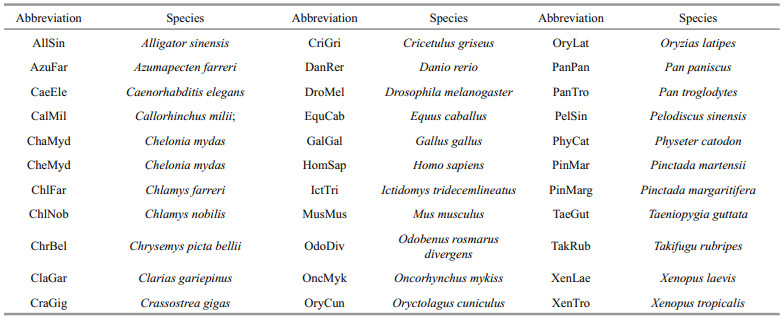
| Fig.3 The phylogenetic tree of the DMRT proteins was constructed by MEGA.6 using the neighbor-joining method (a) and structure model of Pf-DMRT4 protein (b) In a: the number shown at each branch displays the bootstrap value. Each sequence is denoted by the species followed by its subtype. Abbreviations of the species were listed in Supplementary Material 2 and the corresponding GenBank accession numbers of the reference sequences are shown in the phylogenetic tree. In b: yellow indicated zine-fmger-like with DNA binding modules of DM domain. |
A model of the structure of the Pf-DMRT4 was predicted by SWISS Model and the yellow indicated zinc-finger-like with DNA binding modules of DM domain (Fig.3b).
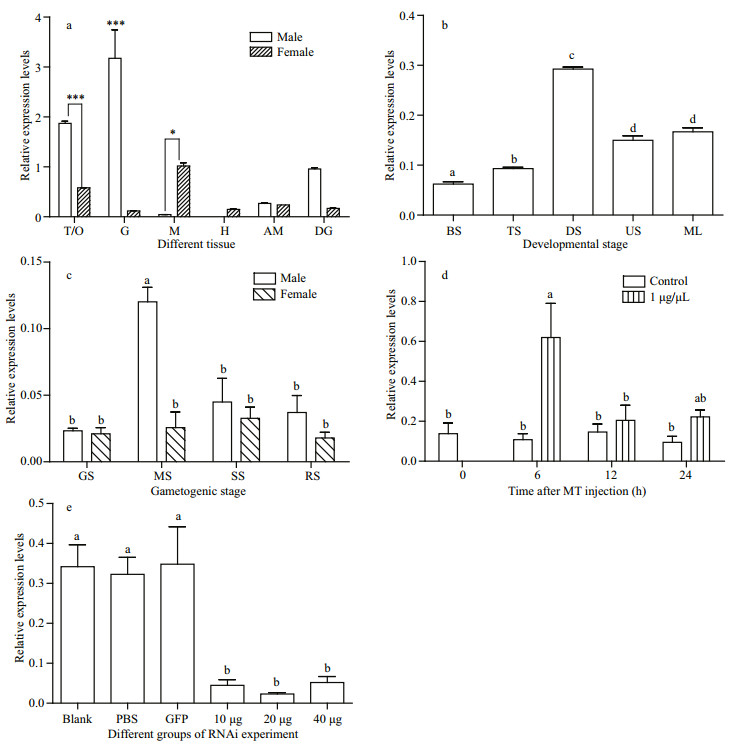
3.2 Pf-Dmrt4 expression pattern in tissuesPf-Dmrt4 mRNA expression levels were detected in all tested tissues by real time qRT-PCR. The expression profiles using df3, dr3 and Sfi, Sr1 primers suggested that mRNA expression reflected Pf-Dmrt4 transcript levels. Pf-Dmrt4mRNA levels were significantly higher in gills and testis, moderate in digestive gland, and low in mantle, adductor muscle and heart tissues. In female tissues, Pf-Dmrt4 mRNA levels were higher in the mantle and ovary compared with other tissues. Pf-Dmrt4 expression levels were higher in testis than in ovaries (Fig. 4a).
|
| Fig.4 Expressions of Pf-Dmrt4 mRNA in tissues of P. fucata (a); expression pattern of Pf-Dmrt4 mRNA in embryonic and larval stages of P. fucata (b); Pf-Dmrt4 mRNA temporal expression in adult gonadic areas of P. fucata along a gametogenetic cycle (c); levels of mRNA expression for Pf-Dmrt4 mRNA in the gonad of P. fucata after MT injection (d); expression levels of Pf-Dmrt4 knocked down by RNAi (e) In a: the mRNA levels were quantified by real-time RT-PCR in: T: testis (male) or O: ovary (female); G: gill; M: mantle; H: heart; AM: adductor muscle; DG: digestive gland. The results are expressed as fold-change. Each bar represents the mean±S.E.M. of three samples. *: P < 0.05; ***: P < 0.001. 18S (AY877529) was used as the reference gene. In b: BS: blastula stage; TS: trochophore stage; DS: D-shaped larva stage; US: umbo larva stage; ML: metamorphosis of larva. Each value represents the mean±SEM (n=3). ***: P < 0.001. In c: GS: growing stage; MS: mature stage; SS: spawning stage; RS: resting stage. Each bar represents the mean±S.E.M. of five samples. Value without common notation (a, b) differ significantly (P < 0.05). In d: each value represents the mean±SEM (n=10). Means not sharing the same superscript are significantly different (P < 0.05). In e: the expression levels of Pf-Dmrt4 mRNA in the testis were measured by real-time quantitative PCR 7 days after injection. Seven oysters were used in each group. Means not sharing the same superscript are significantly different (P < 0.05). |
Pf-Dmrt4 mRNAs were expressed in all embryonic and larval developmental stages examined (Fig. 4b). Pf-Dmrt4 expression was highest in D-shaped larvae.
3.4 Temporal expression patterns of Pf-Dmrt4 during the adult gametogenic cycleThe gametogenic cycle can be divided into four stages: growing stage (active spermatogenesis and growing oocytes), mature stage (mature germ cells and almost no follicular internal), spawning stage (ruptured follicular, spawning mature sperms and oocytes) and resting stage (irregular shape of follicular, contains few spermatogonia and oogonia, degenerate oocytes and sperms can be seen occasionally). Pf-Dmrt4 mRNA expression levels in gonad were detected by real-time qPCR at all adult gametogenic stages (Fig. 4c). Pf-Dmrt4 mRNA levels were significantly higher in the mature stage compared with the other three stages in the male gametogenic cycle. However, there were no significant differences in Pf-Dmrt4 expression among the different stages of the female gametogenic cycle.
3.5 Effects of MT injection on Pf-Dmrt4 expressionPf-Dmrt4 mRNA expression in the gonads during a gametogenic cycle, measured by quantitative polymerase chain reaction, was maximal in mature male testis. Therefore, mature male testis were used in the MT injection experiment.
Pf-Dmrt4 mRNA levels in the testis were significantly increased 6-h post-injection with 1 μg/ μL of MT (Fig. 4d), being almost 3- fold higher than in control groups, respectively.
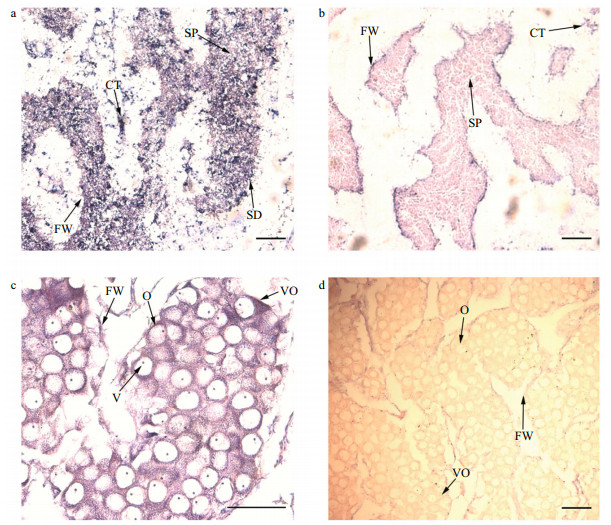
3.6 Localization of Pf-Dmrt4 transcripts in the gonads of P. fucataWe determined the cellular localizations of Pf-Dmrt4 transcripts in mature P. fucata by in situ hybridization. Pf-Dmrt4 mRNA in the male gonads was detected in the cytoplasm of spermatozoa and to a lesser extent in spermatids (Fig. 5a). In the female gonads (Fig. 5c), cytoplasmic mRNA staining was observed in oocytes and vitellogenic oocytes. Negative controls with the sense riboprobe sometimes produced a faint, insignificant signal (Fig. 5b, d).
|
| Fig.5 Pf-Dmrt4 mRNA cellular expression patterns by in-situ hybridization in the gonadic area during the mature stage In males, Pf-Dmrt4 mRNA expression is found in spermatozoa and to a lesser extent in spermatid (a). In females, cytoplasmic staining is observed in oocytes and vitellogenic oocytes (c). Whatever the sex, a somatic staining cannot be excluded. No staining is observed with the sense probe (b, d). SP: spermatozoa; CT: connected tissue; N: nucleus; O: oocyte; VO: vitellogenic oocytes; FW: follicular wall. Bars: 200 μm. |
Pf-Dmrt4 expression levels were decreased by Pf-Dmrt4 dsRNA (10, 20, and 40 μg) respectively (Fig. 4e). Expression levels in the groups injected with 20 μg Pf-Dmrt4 dsRNA were suppressed to approximately 70% and 50% of those in the PBS- and GFP-dsRNA-injected groups, respectively.
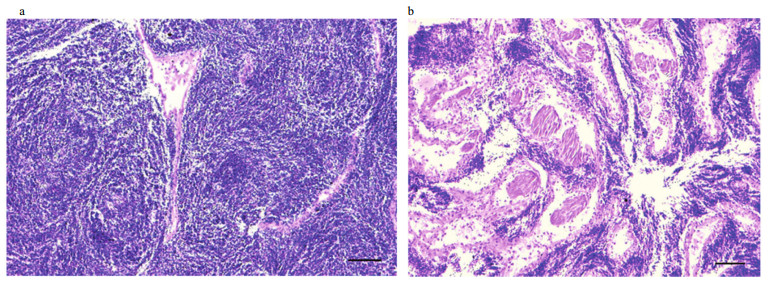
Male gonads of pearl oysters in each injection group were observed under an optical microscope. The gonad structure was normal (mature stage) in the PBS-injected groups (Fig. 6a), while Pf-Dmrt4- dsRNA-injected groups showed spawning-stage male gonads (Fig. 6b), with ruptured follicles and released spermatozoa.
|
| Fig.6 Light microscope images of the testis of the pearl oysters injected with PBS (a), 20 μg of Pf-Dmrt4 (b) dsRNA in the magnification of 100, respectively Bars: 200 μm. |
The Dmrt-family gene Pf-Dmrt4 from the gonads of P. fucata was cloned. The deduced aa sequence of Pf-Dmrt4 is serine- and proline-rich, includes a DM domain characteristic of the DMRT protein family, a conserved DMA domain near the C-terminus, and a short seven-aa motif (KSAFSPI). The DM domain contains the putative NLS (KGHKR) located in the zinc module, consisting of intertwined CCHC and HCCC Zn2+-binding sites. The NLS is characterized as a transcription regulator of the DMRT family and plays a part in DNA-binding and nuclear import (Zhang et al., 2001; Ying et al., 2007). The DMA domain is conserved in the branch of DMRT3, DMRT4 and DMRT5 (Guo et al., 2004). Wen et al. (2009) assumed that the DMA domain may act as a motif to bind other transcriptional co-factors and can probably be essential for the role of DMRT4 to regulate other genes in transcriptional level.
Yu et al. (2009) previously cloned PinMarDmrt5 from P. martensii, which is largely identical to Pf-Dmrt4. However, the deduced PinMarDmrt5 product of 206 aa lacks the DMA domain and the short conserved motif. Sequence alignment of the two genes revealed that PinMarDmrt5 lacked four nucleotides causing a frame shift. The subsequent nucleotide sequence resulted in a stop codon, thus explaining the short product with no DMA domain or conserved motif. P. fucata DMRT4 displayed higher homologies with human DMRT4 and DMRT5 than with human DMRT1-3.
The availability of the P. fucata genome (Takeuchi et al., 2012; Du et al., 2017) provide an opportunity for a comprehensive study of sex-determining pathways in this species. In the genome of the P. fucata, a dmrtA2-like gene was investigated. While pacific oyster genome encodes three DM domain containing genes: Cg19568, Cg01830, and Cg15952 (named CgDMl) (Zhang et al., 2014). They found that high expression in testis supports a possible role for Cg19568 (named CgDsx) in determining or promoting male-specific development. It contains a DM domain showing closest homology (45%, E-value=9e-13) to Dsx isoform A found in D. melanogaster. Pmargdmrt2 sequence was seen to share the highest amino acid identity with P. martensii dmrt2 and zebrafish Dmrt2 (100% and 95%, respectively), and pmargdmrt with P. martensii dmrt2 and mouse Dmrt4 (59% and 58%, respectively). The other two DM domain genes both show the highest homology to DmrtA2 from many species. Phylogenetic analysis showed that Pf-Dmrt4 was grouped with DMRT4 clusters and more closely with human DMRT4. Pf-Dmrt4 is a DM domain gene and a close relative of CgDMl. Volff et al. (2003) proposed a simpler nomenclature (Dmrt1- 8) based on human genes, with no suggestions of structural or phylogenetic relationships between different genes. We therefore named it as Pf-Dmrt4.
4.2 Potential function of Pf-Dmrt4It was reported that ancestral DM proteins possessed a DMA domain (Miller et al., 2003; Volff et al., 2003), and the presence of a DMA domain in PfDMRT4 reinforces the hypothesis. Pf-DMRT4 as an ancestral DM factor, could be engaged in a number of biological processes, including functions associated with the ancestral structure, whereas evolution has led to diverse DM factors.
Pf-Dmrt4 was expressed in all the tested adult tissues, which is similar to DMl in C. gigas (Naimi et al., 2009), Cf-dmrt4-like in Ch. farreri (Feng et al., 2010), and related genes in other species with different expression patterns (Kim et al., 2003; Guo et al., 2004). Differences in DMRT4 expression among species might reflect variations in gene function. In Takifugu rubripes, Dmrt4 is expressed in the spleen, suggesting an involvement in the immune system (Yamaguchi et al., 2006). Additionally, Pf-Dmrt4 mRNA expression levels were highest in male gills, as for dmrt5 in Ch. nobilis (Shi et al., 2014). While, dmrt5 was expressed highest in gills but no sexual difference has been detected in Ch. farreri (Feng et al., 2010) and in C. gigas (Naimi et al., 2009). These findings suggested that Pf-Dmrt4 is involved in the respiratory system.
Pf-Dmrt4 mRNA was also expressed widely in embryonic and larval stages. Its detection at an early stage resembled the situations in Ch. farreri (Feng et al., 2010), Ch. nobilis (Shi et al., 2014), and some other vertebrates such as medaka (Winkler et al., 2004). Cf-dmrt4-like expression was ubiquitous during embryonic development and peaked at the 8-16-cell stage (Feng et al., 2010), while Ch. nobilis dmrt5 expression peaked at the blastula stage (Shi et al., 2014). Pf-Dmrt4 mRNA expression peaked at the D-shaped larva stage, and this stage is a crucial period to build the larva structure, indicating that Pf-Dmrt4 may play a role in early embryonic development, especially in ontogenesis of larva.
Pf-Dmrt4 may also be involved in gonadal development in the pearl oyster. It was expressed in both sexes, but at higher levels in the testis than in the ovary, as reported for Dmrt4 in Ch. farreri (Feng et al., 2010), DMl in C. gigas (Naimi et al., 2009), medaka (Winkler et al., 2004), and the Japanese pufferfish (Yamaguchi et al., 2006). Pf-Dmrt4 expression levels increased significantly during the mature stage in the testis, but there were no significant differences in mRNA expression in the female gonad, consistent with an increase in Cg-DMl in males at the mature stage (Naimi et al., 2009). The number of spermatogonia increase in the mature stage (stage Ⅲ), as a result not only of their proliferation but also of an increase in gonad volume from 5%-40% of the visceral mass in less-mature animals to 60% at this stage (Fabioux et al., 2004). This indicates that Pf-Dmrt4 may be involved in regulating male gonadal development in P. fucata.
In situ hybridization of the testis revealed that Pf-Dmrt4 was expressed in the cytoplasm of spermatozoa and spermatids, though it was impossible to differentiate between germ cell and somatic cell limits at this microscopic level. In C. gigas, Cg-DMl was observed in the cytoplasm of spermatogonia and/or the surrounding somatic cells (Naimi et al., 2009), in agreement with the expression of Dmrt5 in testicular germ cells in zebrafish (Guo et al., 2004). Different expression patterns exist in different species (Raymond et al., 2000; Winkler et al., 2004). Variations in Pf-Dmrt4 localization suggest that it may play divergent roles in the gonad, including involvement in the proliferation of spermatozoa. Knockdown of Pf-Dmrt4 by RNA interference caused morphological changes in the male gonad, indicating that Pf-Dmrt4 might be involved in maintaining the structure of the male gonad in P. fucata.
Sex steroids play an important role in bivalve reproduction and can promote gamete development. Estrogen injection can affect gametogenesis and gametogenesis-related metabolic pathways in invertebrates (Wang and Croll, 2003; Croll and Wang, 2007). MT significantly upregulated Pf-Dmrt4 expression levels in mature-stage male gonads. Though the effects of sex steroid hormones on Dmrt4 or Dmrt5 are largely unknown, Shi et al. (2014) showed that Dmrt5 mRNA in Ch. nobilis male gonads was significantly increased by injection with three different concentrations of MT. Similar findings were observed for Oreochromis aureus Dmrt4, implying that Dmrt4 might be a direct downstream gene under the control of steroids (Cao et al., 2010), and that Pf-Dmrt4 might play a similar role.
5 CONCLUSIONPf-Dmrt4 is involved in the sex-differentiation pathway in the pearl oyster P. fucata. Pf-Dmrt4 is involved in a wide range of biological processes, including cell proliferation and/or regulating male gonadal development and maintaining the structure of the mature testis in P. fucata. More studies are needed to clarify the precise physiological functions of Pf-Dmrt4.
6 DATA AVALABILITY STATEMENTSequence data that support the findings of this study have been deposited in National Center for Biotechnology Information (GenBank) with the primary accession code KM272582.
|
Aoyama S, Shibata K, Tokunaga S, Takase M, Matsui K, Nakamura M. 2003. Expression of Dmrt1 protein in developing and in sex-reversed gonads of amphibians. Cytogenetic and Genome Research, 101(3-4): 295-301.
DOI:10.1159/000074352 |
Burtis K C, Baker B S. 1989. Drosophila doublesex gene controls somatic sexual differentiation by producing alternatively spliced mRNAs encoding related sexspecific polypeptides. Cell, 56(6): 997-1 010.
DOI:10.1016/0092-8674(89)90633-8 |
Cao J L, Chen J J, Wu T T, Gan X, Luo Y J. 2010. Molecular cloning and sexually dimorphic expression of DMRT4 gene in Oreochromis aureus. Molecular Biology Reports, 37(6): 2 781-2 788.
DOI:10.1007/s11033-009-9820-z |
Crémazy F, Berta P, Girard F. 2001. Genome-wide analysis of Sox genes in Drosophila melanogaster. Mechanisms of Development, 109(2): 371-375.
DOI:10.1016/S0925-4773(01)00529-9 |
Croll R P, Wang C D. 2007. Possible roles of sex steroids in the control of reproduction in bivalve molluscs. Aquaculture, 272(1-4): 76-86.
DOI:10.1016/j.aquaculture.2007.06.031 |
Du X D, Fan G Y, Jiao Y, Zhang H, Guo X M, Huang R L, Zheng Z, Bian C, Deng Y W, Wang Q H, Wang Z D, Liang X M, Liang H Y, Shi C C, Zhao X X, Sun F M, Hao R J, Bai J, Liu J L, Chen W B, Liang J L, Liu W Q, Xu Z, Shi Q, Xu X, Zhang G F, Liu X. 2017. The pearl oyster Pinctada fucata martensii genome and multi-omic analyses provide insights into biomineralization. GigaScience, 6(8): 1-12.
|
Fabioux C, Pouvreau S, Le Roux F, Huvet A. 2004. The oyster vasa-like gene: a specific marker of the germline in Crassostrea gigas. Biochemical and Biophysical Research Communications, 315(4): 897-904.
DOI:10.1016/j.bbrc.2004.01.145 |
Feng Z F, Shao M Y, Sun D P, Zhang Z F. 2010. Cloning, characterization and expression analysis of Cf-dmrt4-like gene in Chlamys farreri. Journal of Fishery Sciences of China, 17(5): 930-940.
(in Chinese with English abstract) |
Guo Y Q, Li Q, Gao S, Zhou X, He Y, Shang X, Cheng H H, Zhou R J. 2004. Molecular cloning, characterization, and expression in brain and gonad of Dmrt5 of zebrafish. Biochemical and Biophysical Research Communications, 324(2): 569-575.
DOI:10.1016/j.bbrc.2004.09.085 |
Hodgkin J. 2002. The remarkable ubiquity of DM domain factors as regulators of sexual phenotype: ancestry or aptitude?. Genes & Development, 16(18): 2 322-2 326.
|
Hong C S, Park B Y, Saint-Jeannet J P. 2007. The function of Dmrt genes in vertebrate development: it is not just about sex. Developmental Biology, 310(1): 1-9.
DOI:10.1016/j.ydbio.2007.07.035 |
Klinbunga S, Amparyup P, Khamnamtong B, Hirono I, Aoki T, Jarayabhand P. 2009. Isolation and characterization of testis-specific DMRT1 in the tropical abalone (Haliotis asinina). Biochemical Genetics, 47(1-2): 66-79.
DOI:10.1007/s10528-008-9207-1 |
Leveugle M, Prat K, Popovici C, Birnbaum D, Coulier F. 2004. Phylogenetic analysis of Ciona intestinalis gene superfamilies supports the hypothesis of successive gene expansions. Journal of Molecular Evolution, 58(2): 168-181.
DOI:10.1007/s00239-003-2538-y |
Marı́n I, Baker B S. 1998. The evolutionary dynamics of sex determination. Science, 281(5385): 1 990-1 994.
DOI:10.1126/science.281.5385.1990 |
Miller S W, Hayward D C, Bunch T A, Miller D J, Ball E E, Bardwell V J, Zarkower D, Brower D L. 2003. A DM domain protein from a coral, Acropora millepora, homologous to proteins important for sex determination. Evolution & Development, 5(3): 251-258.
|
Miyashita T, Hanashita T, Toriyama M, Takagi R, Akashika T, Higashikubo N. 2008. Gene cloning and biochemical characterization of the BMP-2 of Pinctada fucata. Bioscience, Biotechnology, and Biochemistry, 72(1): 37-47.
DOI:10.1271/bbb.70302 |
Naimi A, Martinez A S, Specq M L, Mrac A, Diss B, Mathieu M, Sourdaine P. 2009. Identification and expression of a factor of the DM family in the oyster Crassostrea gigas. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology, 152(2): 189-196.
|
Ottolenghi C, Fellous M, Barbieri M, McElreavey K. 2002. Novel paralogy relations among human chromosomes support a link between the phylogeny of doublesex-related genes and the evolution of sex determination. Genomics, 79(3): 333-343.
DOI:10.1006/geno.2002.6711 |
Peng Q L, Pu Y G, Cheng Z H, Nie L W. 2005. Sequence analysis of three Dmrt genes in Macrobrachium rosenbergii. Journal of Fishery Sciences of China, 12(1): 5-9.
(in Chinese with English abstract) |
Raymond C S, Kettlewell J R, Hirsch B, Bardwell V J, Zarkower D. 1999. Expression of Dmrt1 in the genital ridge of mouse and chicken embryos suggests a role in vertebrate sexual development. Developmental Biology, 215(2): 208-220.
DOI:10.1006/dbio.1999.9461 |
Raymond C S, Murphy M W, O'Sullivan M G, Bardwell V J, Zarkower D. 2000. Dmrt1, a gene related to worm and fly sexual regulators, is required for mammalian testis differentiation. Genes & Development, 14(20): 2 587-2 595.
|
Shen X Y, Cui J Z, Yang G P, Gong Q L, Gu Q Q. 2007. Expression detection of DMRTs and two sox9 genes in Takifugu rubripes (Tetraodontidae, Vertebrata). Journal of Ocean University of China, 6(2): 182-186.
DOI:10.1007/s11802-007-0182-7 |
Shen M M, Hodgkin J. 1988. Mab-3, a gene required for sexspecific yolk protein expression and a male-specific lineage in C. elegans. Cell, 54(7): 1 019-1 031.
DOI:10.1016/0092-8674(88)90117-1 |
Shi Y, Wang Q, He M X. 2014. Molecular identification of dmrt2 and dmrt5 and effect of sex steroids on their expressions in Chlamys nobilis. Aquaculture, 426-427(1): 21-30.
|
Suzuki M, Saruwatari K, Kogure T, Yamamoto Y, Nishimura T, Kato T, Nagasawa H. 2009. An acidic matrix protein, Pif, is a key macromolecule for nacre formation. Science, 325(5946): 1 388-1 390.
DOI:10.1126/science.1173793 |
Takeuchi T, Kawashima T, Koyanagi R, Gyoja F, Tanaka M, Ikuta T, Shoguchi E, Fujiwara M, Shinzato C, Hisata K, Fujie M, Usami T, Nagai K, Maeyama K, Okamoto K, Aoki H, Ishikawa T, Masaoka T, Fujiwara A, Endo K, Endo H, Nagasawa H, Kinoshita S, Asakawa S, Watabe S, Satoh N. 2012. Draft genome of the pearl oyster Pinctada fucata: a platform for understanding bivalve biology. DNA Research, 19(2): 117-130.
DOI:10.1093/dnares/dss005 |
Teaniniuraitemoana V, Huvet A, Levy P, Klopp C, Lhuillier E, Gaertner-Mazouni N, Gueguen Y, Le Moullac G. 2014. Gonad transcriptome analysis of pearl oyster Pinctada margaritifera: identification of potential sex differentiation and sex determining genes. BMC Genomics, 15(1): 491.
DOI:10.1186/1471-2164-15-491 |
Volff J N, Zarkower D, Bardwell V J, Schartl M. 2003. Evolutionary dynamics of the DM domain gene family in metazoans. Journal of Molecular Evolution, 57(S1): S241-S249.
|
Wang C, Croll R P. 2003. Effects of sex steroids on in vitro gamete release in the sea scallop, Placopecten magellanicus. Invertebrate Reproduction & Development, 44(2-3): 89-100.
|
Wen A Y, You F, Tan X G, Sun P, Ni J, Zhang Y Q, Xu D D, Wu Z H, Xu Y L, Zhang P J. 2009. Expression pattern of dmrt4 from olive flounder (Paralichthys olivaceus) in adult gonads and during embryogenesis. Fish Physiology and Biochemistry, 35(3): 421-433.
DOI:10.1007/s10695-008-9267-5 |
Yamaguchi A, Lee K H, Fujimoto H, Kadomura K, Yasumoto S, Matsuyama M. 2006. Expression of the DMRT gene and its roles in early gonadal development of the Japanese pufferfish Takifugu rubripes. Comparative Biochemistry and Physiology Part D: Genomics and Proteomics, 1(1): 59-68.
DOI:10.1016/j.cbd.2005.08.003 |
Ying M, Chen B, Tian Y H, Hou Y, Li Q, Shang X, Sun J H, Cheng H H, Zhou R J. 2007. Nuclear import of human sexual regulator DMRT1 is mediated by importin-β. Biochimica et Biophysica Acta Molecular Cell Resaerch, 1773(6): 804-813.
DOI:10.1016/j.bbamcr.2007.03.006 |
Yu F F, Gui J F, Zhou L, Wang M F, Yu X Y. 2009. Cloning and expression characterization of Dmrt5 in Pinctada martensii. Acta Hydrobiologica Sinica, 33(5): 844-850.
(in Chinese with English abstract) DOI:10.3724/SP.J.1035.2009.50844 |
Yu F F, Wang M F, Zhou L, Gui J F, Yu X Y. 2011. Molecular cloning and expression characterization of DMRT2 in Akoya pearl oysters, Pinctada martensii. Journal of Shellfish Research, 30(2): 247-254.
DOI:10.2983/035.030.0208 |
Yu F F, Yu X Y, Wang M F, Zhou L, Gui J F. 2007. Sex reversal phenomena in bivalves and its mechanism. Acta Hydrobiologica Sinica, 31(4): 576-580.
(in Chinese with English abstract) |
Zhang L X, Hua Z C, Ren J G, Meng A M. 2001. The nuclear localization signal of zebrafish terra is located within the DM domain. FEBS Letters, 503(1): 25-29.
DOI:10.1016/S0014-5793(01)02677-1 |
Zhang N, Xu F, Guo X M. 2014. Genomic analysis of the pacific oyster (Crassostrea gigas) reveals possible conservation of vertebrate sex determination in a mollusc. Genes, Genomes, Genetics, 4(11): 2 207-2 217.
|
 2018, Vol. 36
2018, Vol. 36