Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZHOU Liqing, WANG Xuemei, WU Biao, SUN Xiujun, ZHAO Qing, ZHANG Gaowei, LIU Zhihong, YANG Aiguo
- Karyological analysis of the sea cicada Blepharipoda liberate Shen from the Rizhao intertidal zone, China
- Journal of Oceanology and Limnology, 37(1): 169-175
- http://dx.doi.org/10.1007/s00343-019-7320-0
Article History
- Received Nov. 27, 2017
- accepted in principle Feb. 5, 2018
- accepted for publication Mar. 19, 2018
2 Rizhao Ocean & Fisheries Research Institute, Rizhao 276826, China;
3 Laboratory for Marine Fisheries Science and Food Production Processes, Qingdao National Laboratory for Marine Science and Technology, Qingdao 266071, China
Blepharipoda liberate Shen is a commercially valuable seafood species that is highly popular in Shandong Province, China. Blepharipoda liberate cleans intertidal sand, which is ecologically significant. However, there have been few biological studies on B. liberate in recent years. Hou et al. (2017) investigated fatty acid, amino acid, and cholesterol levels in wild B. liberate. Ding et al. (2016) studied the morphological characteristics and genetic diversity of B. liberate and Lopoomastix japonica, basing their studies partly on COⅠ gene. Using a particular COI gene segment for DNA barcoding, Liu et al. (2016) identified the geographical grouping of the germplasm of B. liberate Rizhao. Wang (2015) studied the external characteristics of B. liberate and reported that it is unisexual, and there are several morphological differences between females and males. The genital pores of female B. liberate are located on the ventral base of pereopod Ⅱ, whereas in male B. liberate they are located on the ventral base of pereopod Ⅳ. Moreover, females have four pairs of endopodites on the abdomen, and males have none. To elucidate the variation of species composition, biomass, and size distribution of the fishery resources in the Uljin marine ranching area of Korea, Yoon et al. (2014) investigated four stations between 2009 and 2010, based on dredge sampling, and the results showed that B. liberate (37 714 ind./km2, 1.5%) was one of the dominant species. There have been no subsequent studies on the karyotype of B. liberate.
Externally, B. liberate resembles neither shrimps nor crabs, because pereopod Ⅳ is located in the abdomen (Wang, 2015), B. liberate is known as a sea cicada. Chromosomes are the carriers of genetic material. However, a lack of information about the chromosomes found in B. liberate has discouraged work on species identification and genetic breeding.
In the present study, we characterized the karyotypes of B. liberate in two different developmental stages by analyzing mitotic chromosomal plates obtained from larvae, and from adult females subsequent to egg-laying.
2 MATERIAL AND METHODTo obtain a larger number of metaphases and sharper centromere localization, we acquired phase Ⅲ zoea-larvae (1 cm in length, Fig. 1) and adult female B. liberate subsequent to egg-layings (Fig. 2) from the Rizhao Ocean and Fisheries Research Institute. To ensure that gender judgments were accurate, and referring to the results observed by Wang (2015), we identified four pairs endopodites on the abdomen of each adult female (Fig. 3). Approximately thirty zoea-larvae and three adult female B. liberate, which had laid eggs, were treated with 0.04% colchicine for 30 min. Whole larvae, or the gills or ovaries of adults were used for the chromosomal investigations. The tissues were subjected to hypotonic treatment in 0.075 mol/L potassium chloride for 50 min at room temperature. They were then transferred to pre-cooling Carnoy's fixative (3:1 methanol/acetic acid) with three changes at intervals of 15 min. The metaphase chromosome slides were prepared from the fixed tissue by dissociation using 50% acetic acid, and three drops of dissociated suspension were spattered onto each clean hot slide(at approximately 65℃). After the slides had been air-dried, they were stained with 10% Giemsa solution in phosphate buffer at pH 6.8 for 30 min, and the dye solution was rinsed with distilled water.

|
| Fig.1 Lateral view of phase Ⅲ zoea-larvae of Blepharipoda liberate Scale bar=200 μm. |

|
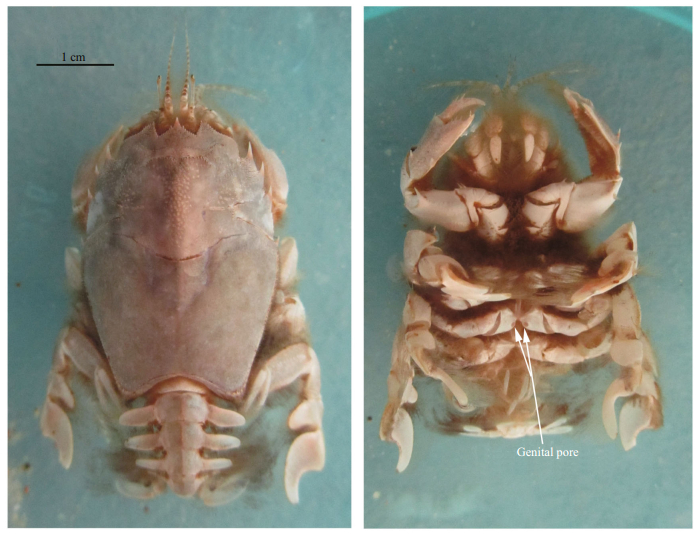
| Fig.2 Dorsal and ventral views of Blepharipoda liberate following egg-laying The arrows indicates the genital pore. Scale bar=1 cm. |
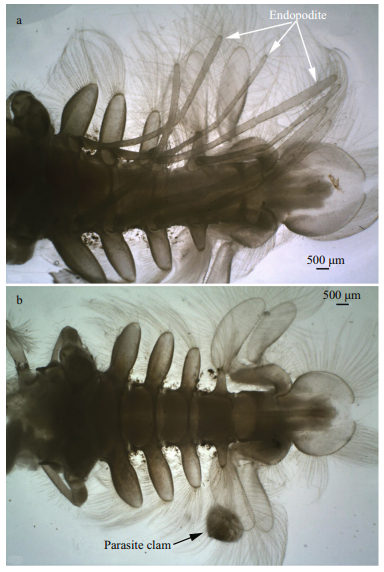
|
| Fig.3 Ventral views of the abdomens of female and male Blepharipoda liberate a. adult female abdomen; b. adult male abdomen. The arrows indicate the endopodites and a parasite clam respectively. Scale bar=500 μm. |
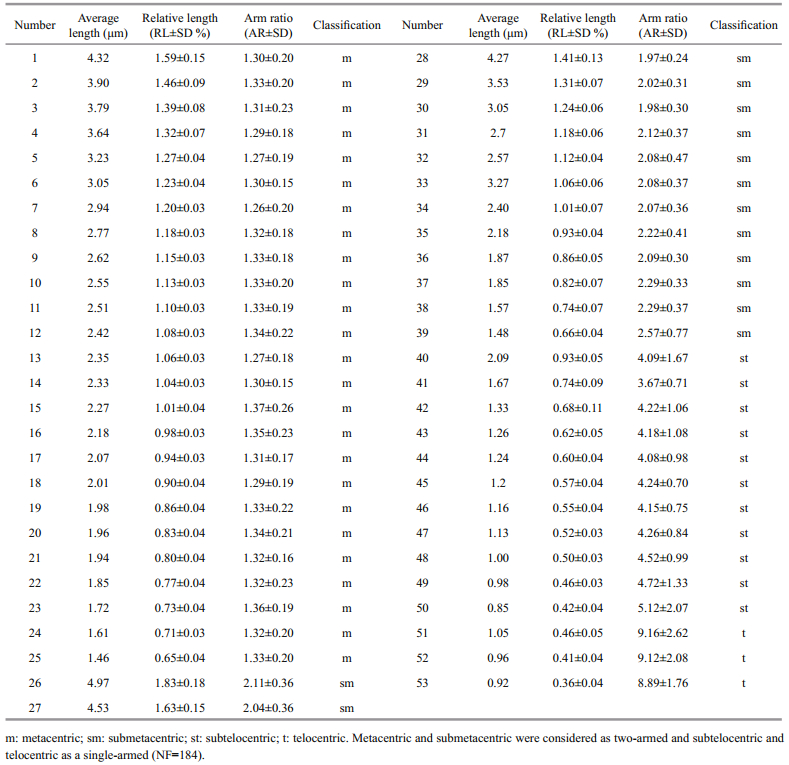
Photomicrographs of the thoroughly spread metaphase plates were obtained using a Leica Microsystem (TYPE DM4000B, Made in Germany). A representative number of metaphases (20–50) from each specimen were analyzed to determine the mode number of the chromosomes. The morphometry of the chromosome pictures was achieved using Adobe Photoshop CS5 photographic software. For each chromosome the average lengths of the short and long arms, the total length, the relative length (RL; the length of each chromosome relative to the sum of the total length of the chromosomes, expressed as a percentage), and the arm ratio (AR; the ratio of the lengths of the long and short arms) were measured and calculated. The chromosome pairs were classified according to the procedure reported by Levan et al. (1964) into metacentric, submetacentric, subtelocentric, and telocentric, with AR ranges of 1-1.7, 1.7-3, 3-7 and 7-∞, respectively.
3 RESULTA total of 218 well-spread mitotic metaphase chromosome sets were examined to determine the diploid chromosome mode number: 183 were from zoea-larvae cells, 23 were from gill cells, and 12 were from the gonad cells of adult females. The modal diploid number (2n) was 106, the numerical variations in the chromosome numbers ranged from 81 to 113, with a mode of 106. We found it easier to investigate chromosomes from larvae cells than gill or gonad cells of adult females, but larval chromosomes were easily lost during preparation (Table 1). Eleven wellspread metaphase chromosome sets were used for karyotype analysis; the karyogram consisted of 25 metacentric, 14 submetacentric, 11 subtelocentric, and 3 telocentric pairs, and we found no heteromorphosis sex chromosomes. The arm number was 184, and the chromosomes varied in size from 0.85 to 4.97 μm (Table 2). A well-spread metaphase plate, and the karyogram of the 106 chromosomes from a larvae cell are shown in Fig. 4 and Fig. 5, respectively. A well-spread metaphase plate from gill cells is shown in Fig. 6.

|
|
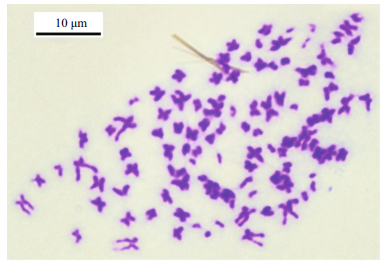
|
| Fig.4 Mitotic metaphase of Blepharipoda liberate Scale indicates 10 μm. |

|
| Fig.5 Karyotype of Blepharipoda liberate with 2n=106 Scale indicates 10 μm. m=metacentric; sm=submetacentric; st=subtelocentric; t=telocentric. |
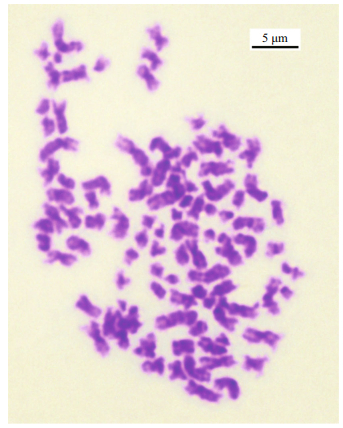
|
| Fig.6 A metaphase plate from Blepharipoda Liberate gill cells Scale indicates 5 μm. |
The arms of the chromosomes in the metaphase plate from the adult female gonads were markedly fluffy (Fig. 7), reminiscent of lampbrush chromosomes, indicating active gene transcription, whereas there were no lampbrush chromosomes in the gill or larvae cells, the chromosome arms of which were relatively dense and round.
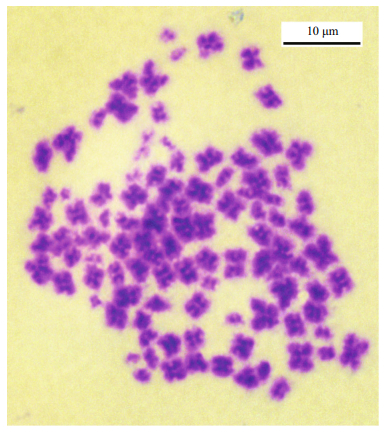
|
| Fig.7 Fluffy chromosomes on a metaphase plate from a Blepharipoda liberate ovary Scale indicates 10 μm. |
There have been few karyological studies of crabs because their chromosomes are numerous, very small, and have indistinct centromeres. Therefore, karyotype analysis is very difficult in many crab species. Lee et al. (2004) reported that it was not possible to distinguish between two species of crab using karyotype analysis because the centromeres of some chromosomes could not be located, and the number of chromosomes could not be used to distinguish between species. Based on our findings, we created a relatively accurate karyotype formula: 2n=50m+28sm+22st+6t, 2n=106, NF=184. However, it was impossible to determine whether the smallest eight pairs of chromosomes were type st or t, and we did not identify the sex chromosome. We aim to carry out restriction site-associated DNA sequencing (RAD-seq) to investigate the sex determination mechanism reported by Mathers et al. (2015).
In the present study, we determined the diploid chromosome number (2n) of B. liberate to be 106. This agrees with the diploid chromosome numbers obtained for Portunus trituberculatus (Zhu et al., 2005), Hippa talpoides, Ranina ranina, and Macrocheira kaempferi (Wang et al., 2005), which are all crabs. The diploid chromosome numbers of most shrimps range from 60 to 90. This implies that B. liberate is more closely relatedly to crabs than shrimps.
Larvae, gills, and gonad tissue can all be used for chromosomal investigation. However, in the present study, we found that the number of metaphase plates from the gills and gonads was low, and the chromosomes did not readily spread out, whereas the larvae tissue yielded more metaphase plates but the chromosomes were easily lost during the dissociation step. Therefore, the dissociation method needs to be improved.
Lampbrush chromosomes (LBCs) are transcriptionally active in various animals during oogenesis, and are also found in the giant singlecelled alga Acetabularia (Gall, 2012). The LBCs of lower vertebrates and invertebrates often appear in the diplotene and dictyate phases, and are probably sites of active gene transcription (Gwatkin, 1977). These chromosomes produce large amounts of RNA for the oocyte (Piprek et al., 2014), and may be encountered during all stages of oocyte development.
We also found LBCs in a metaphase plate from adult female B. liberate gonads, which implies that oogenesis is continuous regardless of ovulation, and the LBCs produced by ovaries are presumed to be homologous in the zygotene phase of oocytes. The microchromosomes lampbrushes has been exquisitely structured, which revealed many fundamentally significant features of the karyotype of Bipes canaliculatus more clearly, and also confirmed that the period of maximum extension of the lampbrush loops occurs well before the start of the major phase of vitellogenesis and oocyte growth (Macgregor and Klosterman, 1979). Moreover, LBCs appear to work synergistically structuring the dynamic growth of the oocyte (Montezol et al., 2018). A comparison of the C-banded karyotypes and the maps of LBCs shows that some of the regions of constitutive heterochromatin seem to correspond to the structures inserted on the LBCs in Triturus species (Schmid et al., 1979). Therefore, the researchers deduced that only the experiments with in situ hybridization of Triturus-DNA could be used to determine whether these structures were really identical to the C-bands. Fluorescent in situ hybridization (FISH) investigations of chicken and Japanese quail LBCs yielded high resolution gene maps (Galkina et al., 2006). Therefore, the lampbrush chromosomes of B. liberate warrant further investigation.
5 CONCLUSIONThe number of chromosomes in B. liberate is 2n=106. Larvae, gills, and ovary tissue can all be used for chromosomal investigation, but larvae are more practical, and ovary tissues are suitable for the preparation of lampbrush chromosomes. Heteromorphosis sex chromosomes have not yet been dioscovered. Based on the chromosome numbers, it appears that B. liberate is more closely related to crabs than to shrimps. The B. liberate karyotype is reported here providing a basis for further comparative cytogenetic studies of the species.
6 DATA AVAILABILITY STATEMENTAll data generated and analyzed during the current study are included in this published article.
7 ACKNOWLEDGEMENTThe authors are grateful to SUN Yuzhong, Director of Rizhao Ocean & Fisheries Research Institute, and to Professor CHEN Siqing and engineer ZHANG Shengnong at the Yellow Sea Fisheries Institute, Chinese Academy of Fishery Sciences for enabling the cytogenetic experiments and chromosome preparations.
Ding P E, Song N, Wang X M, Han Z Q, Gao T X. 2016. The morphological characters and genetic diversity of Blepharipoda liberate Shen and Lophomastix japonica. Acta Hydrobiologica Sinica, 40(6): 1208-1214.
(in Chinese with English abstract) |
Galkina S, Deryusheva S, Fillon V, Vignal A, Crooijmans R, Groenen M, Rodionov A, Gaginskaya E. 2006. FISH on avian lampbrush chromosomes produces higher resolution gene mapping. Genetica, 128(1-3): 241-251.
DOI:10.1007/s10709-005-5776-7 |
Gall J G. 2012. Are lampbrush chromosomes unique to meiotic cells?. Chromosome Research, 20(8): 905-909.
DOI:10.1007/s10577-012-9329-5 |
Gwatkin R B L. 1977. The egg. In: Gwatkin R B L ed.Fertilization Mechanisms in Man and Mammals. Plenum Press, New York, USA. p.5-12.
|
Hou Z H, Song Q W, Cui B. 2017. Analysis of nutrient composition in Blepharipoda liberate Shen captured in the wild. Modern Food Science and Technology, 33(6): 262-270.
(in Chinese with English abstract) |
Lee T H, Naitoh N, Yamazaki F. 2004. Chromosome studies on the mitten crabs Eriocheir japonica and E.sinensis. Fisheries Science, 70(2): 211-214.
DOI:10.1111/j.1444-2906.2003.00793.x |
Levan A, Fredga K, Sandberg A A. 1964. Nomenclature for centromeric position on chromosomes. Hereditas, 52(2): 201-220.
DOI:10.1111/j.1601-5223.1964.tb01953.x |
Liu F M, Li J Z, Liang Q B, Lin S J. 2016. DNA barcoding and molecular evolution of Rizhao Haichan. Journal of Anhui Agriculture Science, 44(3): 133-135.
(in Chinese with English abstract) |
Macgregor H, Klosterman L. 1979. Observations on the cytology of Bipes (Amphisbaenia) with special reference to its lampbrush chromosomes. Chromosoma, 72(1): 67-87.
|
Mathers T C, Hammond R L, Jenner R A, Hänfling B, Atkins J, Gómez A. 2015. Transition in sexual system and sex chromosome evolution in the tadpole shrimp Triops cancriformis. Heredity, 115(1): 37-46.
DOI:10.1038/hdy.2015.10 |
Montezol M, Cassel M, Silva D, Ferreira A, Mehanna M. 2018. Gametogenesis and reproductive dynamics of Rhinella schneideri (Anura:Bufonidae):influence of environmental and anthropogenic factors. Acta Zoologica, 99(1): 93-104.
DOI:10.1111/azo.12195 |
Piprek R P, Kloc M, Kubiak J Z. 2014. Bidder's organ-structure, development and function. The International Journal of Developmental Biology, 58(10-12): 819-827.
DOI:10.1387/ijdb.140147rp |
Schmid M, Olert J, Klett C. 1979. Chromosome banding in Amphibia. Ⅲ. Sex chromosomes in Triturus. Chromosoma, 71(1): 29-55.
|
Wang Q, Kong X Y, Yu S S, Yu Z N. 2005. Advance in study on chromosome of deoapoda. Marine Sciences, 29(6): 60-65.
(in Chinese with English abstract) |
Wang X M. 2015. The morphological characteristics of the Blepharipoda liberate Shen. Progress in Fishery Sciences, 36(1): 74-78.
(in Chinese with English abstract) |
Yoon B S, Park J H, Yoon S C, Yang J H, Sohn M H. 2014. Community structure of fisheries resources caught by dredge in the Uljin marine ranching area, Korea. Korean Journal of Fisheries and Aquatic Sciences, 47(6): 935-944.
DOI:10.5657/KFAS.2014.0935 |
Zhu D F, Wang C L, Li Z Q. 2005. Karyotype analysis on Portunus trituberculatus. Journal of Fisheries of China, 29(5): 649-653.
(in Chinese with English abstract) |
 2019, Vol. 37
2019, Vol. 37


