Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZHANG Wenjing, JIN Weihua, DUAN Delin, ZHANG Quanbin
- Structural analysis and anti-complement activity of polysaccharides extracted from Grateloupia livida (Harv.) Yamada
- Journal of Oceanology and Limnology, 37(3): 806-814
- http://dx.doi.org/10.1007/s00343-019-8125-x
Article History
- Received May. 12, 2018
- accepted in principle Jun. 13, 2018
- accepted for publication Jul. 29, 2018
2 School of Medicine, Zhejiang University, Hangzhou 310058, China;
3 College of Biotechnology and Bioengineering, Zhejiang University of Technology, Hangzhou 310014, China;
4 Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory for Marine Science and Technology, Qingdao 266071, China;
5 State Key Lab of Seaweed Bioactive Substances, Qingdao 266000, China;
6 Center for Ocean Mega-Science, Chinese Academy of Sciences, Qingdao 266071, China
Sulfated polysaccharides from red seaweed possess biological activities including antioxidant, anticoagulant, anti-HIV, anticholinesterase, and antityrosinase activities (Neushul, 1990; Shanmugam and Mody, 2000; Zhang et al., 2003; De Souza et al., 2007; Wang et al., 2007; Lins et al., 2009; Jiang et al., 2013). Sulfated galactans, known as agarans and carrageenans depending on their stereochemistry, coexist in the family Gigartinaceae (Falshaw and Furneaux, 1998). As an important agaran, porphyran has a backbone of alternating 3-linked β-D-galactose and 4-linked α-L-galactose-6-sulfate or 3, 6-anhydro- α-L-galactose units (Duckworth and Yaphe, 1971; Falshaw and Furneaux, 1998; Usov, 1998). Usually, the agaran backbone is interrupted by different O-linked substituents, such as sulfate, methyl, and xylosyl groups (Jiao et al., 2011). Carrageenans are composed of linear repeating disaccharide units with alternating 3-linked β-D-galactopyranosyl and 4-linked α-D-galactopyranosyl units branched with sulfate, pyruvate, or methyl groups. Furthermore, 4-linked units in the form of a 3, 6-anhydride are common (Usov, 1984; Jiao et al., 2011).
Polysaccharide structural elucidation has previously mainly depended on methylation analysis and chemical methods. Although several new analytical procedures have been attempted, including enzymatic hydrolysis, most are tedious and timeconsuming "wet chemistry" methods (Knutsen and Grasdalen, 1992a). Owing to its ability to determine the structures and sulfation patterns of sulfated galactans, nuclear magnetic resonance (NMR) spectroscopy has proven to be more effective at structural determination (Jin et al., 2013a, b; Zhang et al., 2015). However, overlapping signals make NMR spectral analysis difficult (Knutsen and Grasdalen, 1992a, b; Usov, 1998). In the past decade, electrospray ionization collision-induced dissociation tandem mass spectrometry (ESI-CID-MS/MS) has become the primary analytical tool for obtaining information on polysaccharide linkage types, sulfation, and backbone sequences with high sensitivity and selectivity (Zaia et al., 2003; Maslen et al., 2006).
Grateloupia livida (Harv.) Yamada is an epidemic species that grows on lower intertidal rocks in the South China Sea. In recent years, researchers have shown that extracts from the Grateloupia family have several biological activities, including antiangiogenic, antioxidant, antiviral, antiallergic, and anticancer activities (Liu and Pang, 2010; Ngo and Kim, 2013). This study aimed to elucidate the structural features of polysaccharides from Grateloupia livida (Harv.) Yamada by combining ESI-MS with NMR and infrared (IR) spectroscopic analyses. The anticomplement activity of these polysaccharides was also determined for the exploitation of medicines and foods.
2 MATERIAL AND METHOD 2.1 MaterialGrateloupia livida (Harv.) Yamada was collected at Nanao Island, Shantou, Guangdong Province, China. The sample was washed thoroughly with deionized water and dried in air.
2.2 Preparation of polysaccharideBriefly, the dried alga was cut into pieces and the polysaccharide was extracted with hot water at 105℃ for 4 h. The resultant solution was concentrated, dialyzed, and precipitated with ethanol. This crude polysaccharide precipitate was named WGW.
2.3 Preparation of degraded polysaccharideCrude polysaccharide WGW was degraded using 0.5 mol/L sulfuric acid at 90℃ for 3 h. The degraded polysaccharide, named WGWD, was obtained after neutralization, ultrafiltration, concentration, and lyophilization.
2.4 Anion-exchange chromatographyWGWD was separated by anion-exchange chromatography on a DEAE-Bio-Gel agarose FF (5 cm×60 cm) column using water and a linear gradient of NaCl solution (0–2 mol/L) at a flow rate of 10 mL/min. Finally, WGWD was divided into four fractions, named S1, S2, S3, and S4. All fractions were desalted using a Sephadex G-10 Gel column (2.6 cm×100 cm).
2.5 Composition analysisTotal sugar and 3, 6-anhydrogalactose (A) contents were determined according to the methods of DuBois et al. (1956) and Yaphe and Arsenault (1965), respectively, using galactose as a standard. The sulfation level was analyzed using the barium chloride-gelatin method of Kawai et al. (1969). The molecular weight of samples was determined according to a previous study (Jin et al., 2013b).
2.6 Spectroscopic analysisIR spectra were recorded according to a previous study (Jin et al., 2013b). Electrospray ionization mass spectrometry was performed on a Micromass Q-Tof Ultima instrument (Waters, Manchester, UK). Samples were introduced at a flow rate of 5 μL/min in negative ionization mode with a capillary voltage of -3 000 V, cone voltage of -50 V, source temperature of 80℃, and desolvation temperature of 150℃. The collision energy was optimized between 10 and 50 eV. All spectra were analyzed by Xcalibur software.
Polysaccharide sample (50 mg) was co-evaporated twice with deuterium oxide (99.9%) before dissolving in deuterium oxide (99.9%) containing deuterated acetone (0.1 μL). The DEPTQ (distortionless enhancement by polarization transfer including detection of quaternary nuclei) spectrum was recorded using a Bruker AVANCE 600 MHz spectrometer at 25℃.
2.7 Anti-complement activityThe anti-complement activity was determined using the classical pathway according to a previous study (Xu et al., 2007) with modifications. Normal human serum (NHS, obtained from healthy adult donors; 100 μL, diluted 1:10), GVB2+ (veronal buffered saline containing 0.1% gelatin, 0.5 mmol/L Mg2+, and 0.15 mmol/L Ca2+; 200 μL) and sensitized erythrocytes (EA; 200 μL) were added to polysaccharide solutions of various dilutions (100 μL) and incubated at 37℃ for 30 min. The four assays were as follows: (ⅰ) 100% lysis, using EA (200 μL) in water (400 μL); (ⅱ) sample control, using sample (100 μL) in GVB2+ (500 μL); (ⅲ) complement, using 1:10-diluted NHS (100 μL) and EA (200 μL) in GVB2+ (300 μL); and (ⅳ) blank, using EA (200 μL) in GVB2+ (400 μL). These mixtures were later centrifuged and determined at a wavelength of 405 nm. The following equation was used to calculate the inhibition of EA lysis:

Chemical composition analysis showed that WGW contained 12.40% 3, 6-anhydrogalactose, 67.78% total sugars, and 23.58% sulfates. The highperformance gel-permeation chromatography (HPGPC) profile of WGW showed two eluted peaks, indicating that the molecular weights of WGW were 162.4 and 119.8 kDa (Fig.S1).
WGW was degraded using 0.5 mol/L sulfuric acid at 90℃ for 3 h to obtain WGWD. The molecular weights of WGWD were 2.3, 1.7, and 1.1 kDa (Fig. S1). The sulfate and total sugar contents of WGWD were 9.79% and 80.63%, respectively. WGWD was further fractionated by anion-exchange chromatography on a DEAE-Bio-Gel agarose FF column using a gradient elution (Fig.S2). Four fractions were obtained, named S1, S2, S3, and S4, and detected by MS after removing NaCl using a Sephadex G-10 Gel column. S1, S2, S3, and S4 had molecular weights of 1.7, 1.9, 2.1, and 2.9 kDa (Fig. S1), sulfate contents of 1.98%, 5.81%, 10.20%, and 17.25%, and total sugar contents of 90.21%, 84.26%, 75.12%, and 65.99%, respectively.
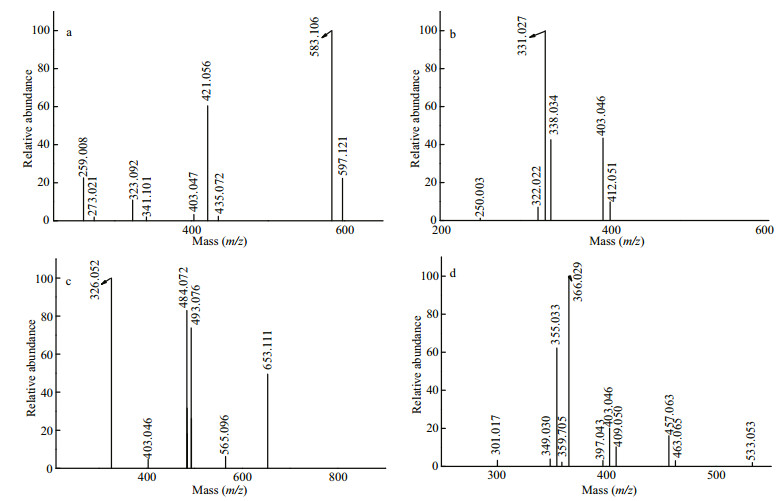
Figure 1a shows the negative ion mode ESI-MS spectrum of S1. Peaks at m/z 259.008,421.056, and 583.106 corresponded to [GalSO3Na-Na]-, [Gal2SO3Na-Na]-, and [Gal3SO3Na-Na]- ions, respectively. Other mass peaks at m/z 273.021,435.072, and 597.122 corresponded to [MeGalSO3NaNa]-, [MeGal2SO3Na-Na]-, and [MeGal3SO3Na-Na]- ions, respectively, while those at m/z 323.092,341.101, and 403.047 corresponded to [GalA-H]- (A: 3, 6-anhydrogalactopyranose residue), [Gal2-H]-, and [GalA(SO3Na)-Na]- ions, respectively.
|
| Fig.1 Negative ion mode ESI-MS spectra of S1 (a), S2 (b), S3 (c), and S4 (d) |
As shown in Figs. 1b, S2 contained ions at m/z 331.027(-2), 338.034(-2), 403.046(-2), 250.003(-2), 322.022(-2), and 412.051(-2) corresponding to [Gal3(SO3Na)2-2Na]2-, [MeGal3(SO3Na)2-2Na]2-, [Gal3A(SO3Na)2-2Na]2-, [Gal2(SO3Na)2-2Na]2-, [Gal2A(SO3Na)2-2Na]2-, and [Gal4(SO3Na)2-2Na]2-, respectively.
As shown in Fig. 1c, the doubly charged ion at m/z 326.052(-2) and singly charged ion at 653.111(-1) corresponded to [Gal3(SO3Na)(PA)-Na]- (PA: pyruvic acid residue), while the doubly charged ions at m/z 484.072(-2) and 493.076(-2) corresponded to [Gal4A(SO3Na)2-2Na]2- and [Gal5(SO3Na)2-2Na]2-, respectively. These were accompanied by [Gal3A(SO3Na)2-2Na]2- (403.046(-2)) and [Gal5A(SO3Na)2-2Na]2- (565.096(-2)).
For S4, the intense signal at m/z 366.029(-2) corresponded to Gal3(SO3Na)2 (PA), while the ions at m/z 355.033(-3) and 533.053(-2) corresponded to Gal5(SO3Na)3 (Fig. 1d). A series of less intense triply charged ions at m/z 349.030,403.046, and 457.063 corresponded to Gal4A(SO3Na)3, Gal5A(SO3Na)3, and Gal6A(SO3Na)3, respectively. Another series of triply charged ions at 301.017,409.050, and 463.065 corresponded to Gal4(SO3Na)3, Gal6(SO3Na)3, and Gal7(SO3Na)3, respectively. S4 also contained MeGal5(SO3Na)3 (359.705(-3)) and Gal4A2(SO3Na)3 (397.043(-3)).
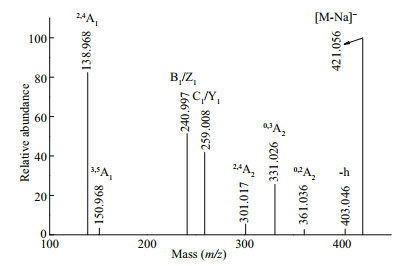
The structural features of the fractions were further analyzed by ESI-CID-MS/MS. The fragmentation pattern of the singly charged ion at m/z 421.056, corresponding to Gal2(SO3Na), is shown in Fig. 2. The fragment ion at m/z 403.046 corresponded to the loss of H2O (-18 Da). The fragment ions at m/z 301.017 (2, 4A2), 331.026 (0, 3A2), and 361.036 (0, 2A2) confirmed that the galactose residues had a major 1→4 linkage accompanied by a minor 1→3 linkage. Furthermore, the presence of an intense signal at 138.968 (2, 4A1) indicated that the sulfate group was located at C4 at the non-reducing end, which was consistent with the IR data. The less intense signal at 150.968 (3, 5A1) indicated that sulfate group was located at C6 at the non-reducing end. In summary, the structure of Gal2(SO3Na) was a mixture of four isomers, namely, Gal(SO3Na)(1→3)Gal, Gal(1→3)Gal(SO3Na), Gal(SO3Na)(1→4)Gal, and Gal(1→4)Gal(SO3Na).
|
| Fig.2 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 421.056 (Gal2SO3Na) |
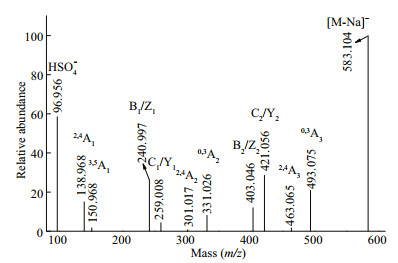
The fragmentation pattern (Fig. 3) of the ion at m/z 583.104 (Gal3(SO3Na)) was similar to that of Gal2SO3Na, suggesting that the fragmentation patterns of single-sulfate galacto-oligosaccharides were similar. The polysaccharides in G. livida (Harv.) Yamada were a mixture of agar and carrageenan, which both had backbones consisting of alternating 3-linked and 4-linked galactopyranose residues. Therefore, we speculated that Gal3SO3Na was a mixture of GalSO3Na(1→3)Gal(1→4)Gal, Gal(1→3) Gal(1→4)GalSO3Na, GalSO3Na(1→4)Gal(1→3) Gal, and Gal (1→4)Gal (1→3)GalSO3Na.
|
| Fig.3 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 583.104 (Gal3SO3Na) |
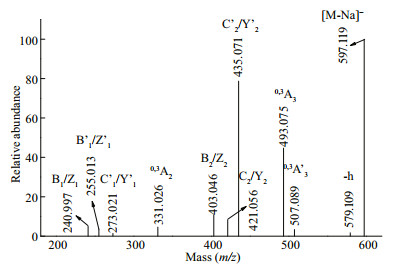
The fragmentation pattern (Fig. 4) of the ion at m/z 597.119 (MeGal3SO3Na) was slightly different from that of Gal3(SO3Na). Differences in the intensities of ions at m/z 331.026 (0, 3A2), 493.075 (0, 3A3), and 507.089 (0, 3A3) indicated two backbones, namely, Gal(1→3)Gal(1→4)Gal and Gal(1→4)Gal(1→3) Gal. Furthermore, the fragmentation ion at m/z 493.075 (0, 3A3) (-104 Da) indicated that the methyl residue was at the reducing end of the 4-linked galactosyl residue. Based on these results, we speculated that the methyl group was positioned at C2. The methyl group in agar was located at C6 in the 3-linked galactopyranose residue and at C2 in the 4-linked galactopyranose residue (Ji, 1997). Furthermore, the presence of a singly charged ion at m/z 331.026 (0, 3A2) confirmed that the methyl group was positioned at C2 in the 4-linked galactopyranose residue. However, methylation at C6 in the 3-linked galactopyranose residue cannot be excluded. Therefore, we concluded that the structures of MeGal3SO3Na were a mixture of isomers Gal(SO3Na) (1→3)Gal(1→4)Gal(2Me), Gal(1→3)Gal(1→4) Gal(2Me)(SO3Na), Gal(SO3Na)(1→4)Gal(2Me) (1→3)Gal, and Gal(1→4)Gal(2Me)(1→3) Gal(SO3Na).
|
| Fig.4 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 597.119 (MeGal3(SO3Na)) |
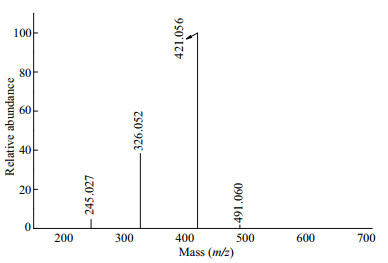
The fragmentation pattern of the doubly charged ion at m/z 326.052(-2), assigned to Gal3(SO3Na)(PA), is shown in Fig. 5. The singly charged ion at m/z 491.060 and a doubly charged ion at m/z 245.027(-2) corresponded to Gal2(SO3Na)(PA) (loss of galactopyranose residue, -162 Da). The ion at m/z 421.056 was derived from the loss of pyruvic acid residue (-70 Da) and galactopyranose residue (-162 Da). The pyruvic acid residue was substituted at C4 and C6 in the 3-linked galactopyranose residue. Therefore, we speculated that Gal3(SO3Na)(PA) was Gal(SO3Na)(1→3)Gal(PA)(1→4)Gal, Gal(1→3) Gal(PA)(1→4)Gal(SO3Na), or Gal(SO3Na)(1→4) Gal(1→3)Gal(PA).
|
| Fig.5 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 326.052(-2) (Gal3(SO3Na)(PA)) |
Figure 6 shows the ESI-CID-MS/MS spectrum of the ion at m/z 484.070(-2) (Gal4A(SO3Na)2). The fragmentation patterns of the ions at m/z 322.022(-2) and 403.046(-2), corresponding to Gal2A(SO3Na)2 and Gal3A(SO3Na)2, were similar to that of Gal4A(SO3Na)2. The characteristic ions at m/z 439.055(-2) (0, 3A4) and 358.031(-2) (0, 3A3) showed that the oligosaccharides contained a backbone with alternating 1, 3-linkages and 1, 4-linkages. According to a previous study (Usov, 1998), 3, 6-anhydrogalactopyranose (residue A) must be 4-linked, suggesting that determining the location of residue A also confirmed the sequence and glycosidic bond of the oligosaccharides. The doubly charged ions at m/z 403.046(-2), 331.026(-2), and 322.021(-2) were derived from the loss of a galactose residue (-162 Da), the loss of galactose (-162 Da) and 3, 6-anhydrogalactopyranose (-144 Da) residues, and the loss of two galactose residues (-324 Da), respectively.

|
| Fig.6 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 484.070(-2) (Gal4A(SO3Na)2) |
According to a previous study (Jin et al., 2013b), a sulfate in the middle of the backbone can easily be lost under acidic conditions. This suggested that the sulfate group was located at a reducing or nonreducing end. Therefore, two backbones were included, as follows: (i) With sulfate groups at the non-reducing end and Gal4A(SO3Na)2 as a mixture of Gal(SO3Na)(1→3)Gal(SO3Na)(1→4)A(1→3) Gal(1→4)Gal or Gal(SO3Na)(1→4)Gal(SO3Na) (1→3)Gal(1→4)A(1→3)Gal isomers; and (ii) with sulfate groups at the reducing end and a mixture of Gal(1→3)Gal(1→4)A(1→3)Gal(SO3Na)(1→4) Gal(SO3Na) or Gal(1→4)A(1→3)Gal(1→4) Gal(SO3Na)(1→3)Gal(SO3Na) isomers. Singly charged ions at m/z 727.142,583.104,583.104, and 421.056 were generated by the loss of a sulfated galactose residue (-243 Da), the loss of a 3, 6-anhydrogalactopyranose residue (-144 Da), the loss of a galactose residue (-162 Da), and the loss of galactose (-162 Da) and 3, 6-anhydrogalactopyranose (-144 Da) residues, respectively. These results indicate that the sulfate groups were located at the reducing and non-reducing ends, which includes Gal(SO3Na)(1→3)Gal(1→4)A(1→3)Gal(1→4) Gal(SO3Na), Gal(SO3Na)(1→4)Gal(1→3)Gal(1→4) A(1→3)Gal(SO3Na), or Gal(SO3Na)(1→4)A(1→3) Gal(1→4)Gal(1→3)Gal(SO3Na) isomers. Therefore, Gal4A(SO3Na)2 might have up to seven isomers.
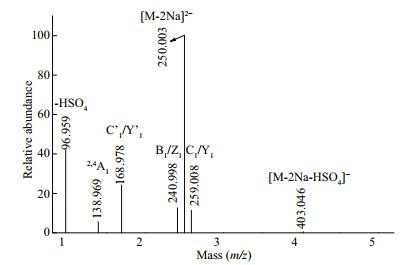
The fragmentation pattern of the doubly charged ion at m/z 250.003(-2) was identified as Gal2(SO3Na)2 in Fig. 7. The doubly charged ion at m/z 168.978(-2) corresponded to Gal(SO3Na)2, indicating that two sulfate groups were substituted at one galactose residue. The ions at m/z 403.046 were derived from the loss of H2SO4 (-98 Da), with the presence of a fragment ion at m/z 96.959 confirming that the sulfate group was easily lost. The ions at m/z 240.998 and 259.008 suggested that sulfate groups were located at the reducing and non-reducing ends. The characteristic ion at m/z 138.969 (2, 4A1) showed that the sulfate group was located at C4 of the reducing end. Therefore, we concluded that Gal2(SO3Na)2 included Gal(SO3Na)2(1→3)Gal, Gal(SO3Na)2(1→4)Gal, Gal(1→3)Gal(SO3Na)2, Gal(1→4)Gal(SO3Na)2, Gal(SO3Na)(1→4)Gal(SO3Na), or Gal(SO3Na)(1→3) Gal(SO3Na).
|
| Fig.7 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 250.003(-2) (Gal2(SO3Na)2) |
The fragmentation patterns of the doubly charged ions at m/z 331.027(-2), 412.051(-2), and 493.075(-2) corresponded to Gal3(SO3Na)2, Gal4(SO3Na)2, and Gal5(SO3Na)2, respectively, and were similar to that of Gal2(SO3Na)2.
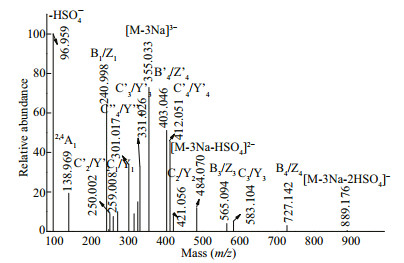
The fragmentation pattern of the triply charged ion at m/z 355.033 (-3) was assigned to Gal5(SO3Na)3 (Fig. 8). Three series of fragment ions were present. The first series included triply charged fragment ions at m/z 325.023(-3) (0, 2A5), 315.020(-3) (2, 4A5), 301.017(-3) (C''4/Y''4), 271.007(-3) (0, 2A5), and 247.001(-3) (C''3/Y''3), which confirmed that three sulfate groups were located at the reducing or nonreducing ends. Therefore, Gal5(SO3Na)3 was a mixture of Gal(SO3Na)(1→3)Gal(SO3Na)(1→4)Gal(SO3Na) (1→3)Gal(1→4)Gal, Gal(SO3Na)(1→4)Gal(SO3Na) (1→3)Gal(SO3Na)(1→4)Gal(1→3)Gal, Gal(1→3) Gal(1→4)Gal(SO3Na)(1→3)Gal(SO3Na)(1→4) Gal(SO3Na), or Gal(1→4)Gal(1→3)Gal(SO3Na) (1→4)Gal(SO3Na)(1→3)Gal(SO3Na) isomers. The second series included a doubly charged fragment ion at m/z 484.070(-2) derived from the loss of H2SO4 (-98 Da). Doubly charged ions at 412.051(-2) (C'4/Y'4), 403.046(-2) (B'4/Z'4), 331.026(-2) (C'3/Y'3), and 250.002(-2) (C'2/Y'2) indicated that two sulfate groups were present at the reducing or non-reducing ends, while the remaining sulfate was randomly located. Therefore, it was concluded that, in addition to the above results, Gal5(SO3Na)3 could also be Gal(SO3Na)(1→3)Gal(SO3Na)(1→4)Gal(1→3) Gal(1→4)Gal(SO3Na), Gal(SO3Na)(1→3)Gal(1→4) Gal(1→3)Gal(SO3Na)(1→4)Gal(SO3Na), Gal(SO3Na) (1→4)Gal(SO3Na)(1→3)Gal(1→4)Gal(1→3)Gal(SO3Na), or Gal(SO3Na)(1→4)Gal(1→3)Gal(1→4) Gal(SO3Na)(1→3)Gal(SO3Na) isomers. The third series included singly charged ions at m/z 889.176 generated by the loss of two H2SO4 groups (-196 Da). Furthermore, singly charged ions at m/z 727.142 (B4/ Z4), 583.104 (C3/Y3), 565.094 (B3/Z3), 421.056 (C2/ Y2), 259.008 (C1/Y1), and 240.998 (B1/Z1) proved that Gal5(SO3Na)3 contained the above eight isomers.
|
| Fig.8 Negative ion mode ESI-CID-MS/MS spectrum of the ion at m/z 355.033(-3) (Gal5(SO3Na)3) |
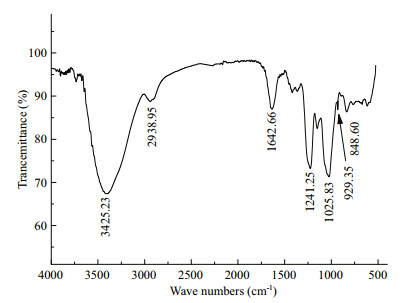
The IR spectrum of WGW (Fig. 9) showed an intense band at approximately 1 240 cm-1, which is characteristic of sulfate esters. A weaker yet discernible signal at approximately 930 cm-1 corresponded to the 3, 6-anhydrogalactosyl units, which was consistent with the MS results. The band at approximately 850 cm-1 indicated that the sulfate group was at the C-4 axial position of galactose. The absence of bands at 830 and 820 cm-1 indicated that there were few or no sulfate groups at C2, C3, and C6.
|
| Fig.9 IR spectrum of WGW |
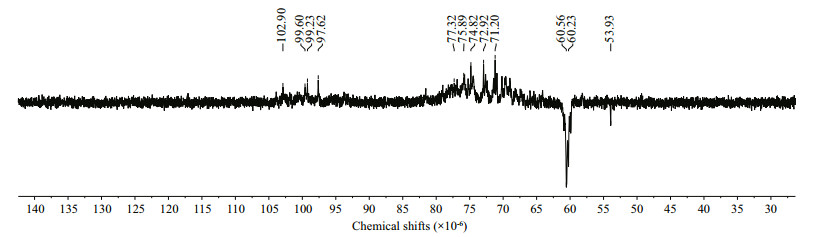
The IR results show that the sulfate in WGW was located at C4. However, 3, 6-anhydrogalactopyranose was 4-linked. Therefore, the sulfate group was located in the β-D-galactose residue. The DEPTQ spectrum of WGW (Fig. 10) was complex, but four recognizable peaks were present at approximately 100×10-6. In combination with previous data (Ji et al., 1997), the polysaccharide structures were proposed as 1, 3-β-DGal (C4S or C6S)(C4S means C4 is sulfated) or 1, 3-β-D-Gal at 102.90×10-6, 1, 4-α-L-Gal (C6S) or 1, 4-α-D-Gal (C2S and C6S) at 99.60×10-6, 1, 4-α-DGal(C6S) at 99.23×10-6, and 1, 4-α-3, 6-anhydro-DGal at 97.62×10-6.
|
| Fig.10 DEPTQ spectrum of WGW |
In conclusion, the structures of polysaccharides from G. livida in this study were analyzed using ESIMS with NMR and IR methods. The high accuracy, sensitivity, selectivity, and speed of MS met the requirements for heteropolysaccharide analysis. However, MS was unable to determine the isomers, which could be resolved by NMR. The polysaccharide from G. livida is an agar–carrageenan intermediate polysaccharide. In this study, this polysaccharide was a mixture of μ-carrageenan, κ-carrageenan, ν-carrageenan, and the agarose precursor. μ-Carrageenan usually has a backbone of alternating 1, 3-linked β-D-galactopyranose residues sulfated at C-4 and 1, 4-linked α-D-galactopyranose residues sulfated at C-6, while κ-carrageenan consists of alternating 1, 3-linked β-D-galactopyranose residues sulfated at C-4 and 1, 4-linked α-D- 3, 6-anhydrogalactopyranose residues. Only trace ν-carrageenan, which is composed of 1, 3-linked β-Dgalactopyranose residues sulfated at C-4 and 1, 4-linked α-D-galactopyranose residues sulfated at C-2 and C-6, was observed. Furthermore, the polysaccharides had a backbone of 1, 3-linked β-Dgalactopyranose and 1, 4-linked α-L-galactopyranose sulfated at C-6, which is the precursor to agarose. Finally, the hydroxy groups in the galactopyranose were found partially substituted by methyl and pyruvic acid acetal (PA) groups.
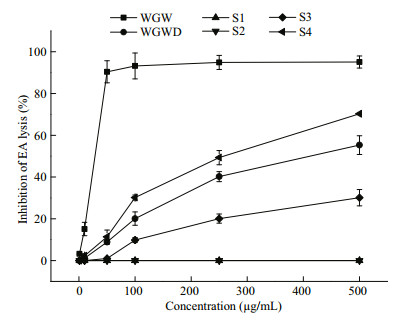
3.2 Anticomplement activityThe complement pathway plays an important part in the immune system, with inappropriate activation causing certain diseases. As shown in Fig. 11, the anticomplement effects of WGW polysaccharides and their derivatives in the classical pathway were examined. The complement group showed 93.56%±4.96% activation of the classical complement pathway. The activities of WGW, WGWD, S3, and S4 were dose-dependent. However, S1 and S2 showed no activity. Although WGW and WGWD had similar chemical compositions, the activities of WGW plateaued at a concentration of 60 μg/mL, while WGWD had a much weaker anti-complement activity, showing an inhibiting rate of 58% at a concentration of 500 μg/mL. The activities were in the order WGW > S4 > WGWD > S3. This indicated that molecular weight and sulfate content were important factors affecting the anti-complement activity, which was consistent with previous studies (Chen et al., 2016; Jin et al., 2016).
|
| Fig.11 Inhibition of classical-pathway-mediated hemolysis of EA in the presence of polysaccharides WGW and WGWD |
Polysaccharide WGW was extracted from G. livida (Harv.) Yamada using hot water. WGW was then degraded by sulfuric acid to obtain WGWD. WGWD was fractionated further by anion-exchange chromatography into four fractions (S1, S2, S3, and S4) and their structural features were elucidated using ESI-MS, ESI-CID-MS/MS, IR, and NMR analyses. WGW was a mixture of μ-carrageenan, κ-carrageenan, ν-carrageenan, and the agarose precursor. The anticomplement activity of WGW and its derivatives against the classical pathway were measured. Native WGW polysaccharides showed higher activity, while the derivatives showed much weaker activity. These data suggested that changes in the molecular weight and sulfate content influenced the anti-complement activities. In summary, polysaccharides from G. livida might be potential drug candidates for anticomplement therapy.
5 DATA AVAILABILITY STATEMENTAll data generated or analyzed in this study are included in this published article.
6 ACKNOWLEDGMENTWe thank Simon Partridge, Ph.D., from Liwen Bianji, Edanz Editing China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Chen M M, Wu J J, Shi S S, Chen Y L, Wang H J, Fan H W, Wang S C. 2016. Structure analysis of a heteropolysaccharide from Taraxacum mongolicum Hand.-Mazz. and anticomplementary activity of its sulfated derivatives. Carbohydrate Polymers, 152: 241-252.
DOI:10.1016/j.carbpol.2016.06.110 |
De Souza M C R, Marques C T, Dore C M G, Da Silva F R F, Rocha H A O, Leite E L. 2007. Antioxidant activities of sulfated polysaccharides from brown and red seaweeds. Journal of Applied Phycology, 19(2): 153-160.
DOI:10.1007/s10811-006-9121-z |
DuBois M, Gilles K A, Hamilton J K, Rebers P A, Smith F. 1956. Colorimetric method for determination of sugars and related substances. Analytical Chemistry, 28(3): 350-356.
DOI:10.1021/ac60111a017 |
Duckworth M, Yaphe W. 1971. The structure of agar:part I.Fractionation of a complex mixture of polysaccharides. Carbohydrate Research, 16(1): 189-197.
DOI:10.1016/S0008-6215(00)86113-3 |
Falshaw R, Furneaux R H. 1998. Structural analysis of carrageenans from the tetrasporic stages of the red algae, Gigartina lanceata and Gigartina chapmanii (Gigartinaceae, Rhodophyta). Carbohydrate Research, 307(3-4): 325-331.
DOI:10.1016/S0008-6215(98)00030-5 |
Ji M H. 1997. Seaweed Chemistry. Science Press, Beijing, China. p.69-70,143-164.
(in Chinese)
|
Jiang Z B, Chen Y C, Yao F, Chen W Z, Zhong S P, Zheng F C, Shi G G. 2013. Antioxidant, antibacterial and antischistosomal activities of extracts from Grateloupia livida (Harv.) Yamada. PLoS One, 8(11): e80413.
DOI:10.1371/journal.pone.0080413 |
Jiao G L, Yu G L, Zhang J Z, Ewart H S. 2011. Chemical structures and bioactivities of sulfated polysaccharides from marine algae. Marine Drugs, 9(2): 196-223.
DOI:10.3390/md9020196 |
Jin W H, Guo Z M, Wang J, Zhang W J, Zhang Q B. 2013a. Structural analysis of sulfated fucan from Saccharina japonica by electrospray ionization tandem mass spectrometry. Carbohydrate Research, 369: 63-67.
DOI:10.1016/j.carres.2012.12.004 |
Jin W H, Zhang W J, Liang H Z, Zhang Q B. 2016. The structure-activity relationship between marine algae polysaccharides and anti-complement activity. Marine Drugs, 14(1): 3.
|
Jin W H, Zhang W J, Wang J, Ren S M, Song N, Zhang Q B. 2013b. Structural analysis of heteropolysaccharide from Saccharina japonica and its derived oligosaccharides. International Journal of Biological Macromolecules, 62: 697-704.
DOI:10.1016/j.ijbiomac.2013.10.017 |
Kawai Y, Seno N, Anno K. 1969. A modified method for chondrosulfatase assay. Analytical Biochemistry, 32(2): 314-321.
|
Knutsen S H, Grasdalen H. 1992a. Analysis of carrageenans by enzymic degradation, gel filtration and 1H NMR spectroscopy. Carbohydrate Polymers, 19(3): 199-210.
DOI:10.1016/0144-8617(92)90162-J |
Knutsen S H, Grasdalen H. 1992b. The use of neocarrabiose oligosaccharides with different length and sulphate substitution as model compounds for 1H-NMR spectroscopy. Carbohydrate Research, 229(2): 233-244.
DOI:10.1016/S0008-6215(00)90573-1 |
Lins K O A L, Bezerra D P, Alves A P N N, Alencar N M N, Lima M W, Torres V M, Farias W R L, Pessoa C, De Moraes M O, Costa-Lotufo L V. 2009. Antitumor properties of a sulfated polysaccharide from the red seaweed Champia feldmannii (Diaz-Pifferer). Journal of Applied Toxicology, 29(1): 20-26.
DOI:10.1002/jat.v29:1 |
Liu F, Pang S J. 2010. Stress tolerance and antioxidant enzymatic activities in the metabolisms of the reactive oxygen species in two intertidal red algae Grateloupia turuturu and Palmaria palmata. Journal of Experimental Marine Biology and Ecology, 382(2): 82-87.
DOI:10.1016/j.jembe.2009.11.005 |
Maslen S, Sadowski P, Adam A, Lilly K, Stephens E. 2006. Differentiation of isomeric N-glycan structures by normalphase liquid chromatography-MALDI-TOF/TOF tandem mass spectrometry. Analytical Chemistry, 78(24): 8491-8498.
DOI:10.1021/ac0614137 |
Neushul M. 1990. Antiviral carbohydrates from marine red algae. Hydrobiologia, 204(1): 99-104.
|
Ngo D H, Kim S K. 2013. Sulfated polysaccharides as bioactive agents from marine algae. International Journal of Biological Macromolecules, 62: 70-75.
DOI:10.1016/j.ijbiomac.2013.08.036 |
Shanmugam M, Mody K H. 2000. Heparinoid-active sulphated polysaccharides from marine algae as potential blood anticoagulant agents. Current Science, 79(12): 1672-1683.
|
Usov A I. 1984. NMR spectroscopy of red seaweed polysaccharides:agars, carrageenans, and xylans. Botanica Marina, 27(5): 189-202.
|
Usov A I. 1998. Structural analysis of red seaweed galactans of agar and carrageenan groups. Food Hydrocolloids, 12(3): 301-308.
DOI:10.1016/S0268-005X(98)00018-6 |
Wang S C, Bligh S W A, Shi S S, Wang Z T, Hu Z B, Crowder J, Branford-White C, Vella C. 2007. Structural features and anti-HIV-1 activity of novel polysaccharides from red algae Grateloupia longifolia and Grateloupia filicina. International Journal of Biological Macromolecules, 41(4): 369-375.
DOI:10.1016/j.ijbiomac.2007.05.008 |
Xu H, Zhang Y Y, Zhang J W, Chen D F. 2007. Isolation and characterization of an anti-complementary polysaccharide D3-S1 from the roots of Bupleurum smithii. International Immunopharmacology, 7(2): 175-182.
DOI:10.1016/j.intimp.2006.09.006 |
Yaphe W, Arsenault G P. 1965. Improved resorcinol reagent for the determination of fructose, and of 3, 6-anhydrogalactose in polysaccharides. Analytical Biochemistry, 13(1): 143-148.
|
Zaia J, Li X Q, Chan S Y, Costello C E. 2003. Tandem mass spectrometric strategies for determination of sulfation positions and uronic acid epimerization in chondroitin sulfate oligosaccharides. Journal of the American Society for Mass Spectrometry, 14(11): 1270-1281.
DOI:10.1016/S1044-0305(03)00541-5 |
Zhang Q B, Yu P Z, Li Z E, Zhang H, Xu Z H, Li P C. 2003. Antioxidant activities of sulfated polysaccharide fractions from Porphyra haitanesis. Journal of Applied Phycology, 15(4): 305-310.
DOI:10.1023/A:1025137728525 |
Zhang W J, Jin W H, Sun D L, Zhao L Y, Wang J, Duan D L, Zhang Q B. 2015. Structural analysis and anti-complement activity of polysaccharides from Kjellmaniella crsaaifolia. Marine Drugs, 13(3): 1360-1374.
DOI:10.3390/md13031360 |
 2019, Vol. 37
2019, Vol. 37


