Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZHANG Wenyan, LIU Jia, DONG Yi, LI Xuegang, XU Cong, XIAO Tian, PAN Hongmiao, WU Long-Fei
- Archaeal community structure in sediments from a seamount in the Mariana Volcanic Arc
- Journal of Oceanology and Limnology, 37(4): 1197-1210
- http://dx.doi.org/10.1007/s00343-019-8044-x
Article History
- Received Mar. 22, 2018
- accepted in principle May. 21, 2018
- accepted for publication Aug. 23, 2018
2 Laboratory for Marine Ecology and Environmental Science, Qingdao National Laboratory for Marine Science and Technology, Qingdao 266071, China;
3 University of Chinese Academy of Sciences, Beijing 100049, China;
4 International Associated Laboratory of Evolution and Development of Magnetotactic Multicellular Organisms (LIA-MagMC), CNRS-CAS, Marseille-Beijing-Qingdao-Sanya;
5 Aix Marseille Univ, CNRS, LCB, Marseille, 31 chemin Joseph Aiguier, F-13402, France;
6 Center for Ocean Mega-Science, Chinese Academy of Sciences, Qingdao 266071, China
Archaea is the third life domain, alongside Bacteria and Eukarya (Woese and Fox, 1977; Woese et al., 1990), whose species were once thought to only occur in extreme environments. However, recent molecular biology tools have revealed that Archaea have a wide ecological distribution, in that they occur in freshwater sediments, soils, and other habitats, and they also have important roles in global biogeochemical cycles (Fuhrman et al., 1992; Auguet et al., 2010). In marine environments, Archaea account for nearly 30% of microbial plankton (DeLong, 1992; Karner et al., 2001; Bano et al., 2004), and are one of the most dominant groups of prokaryotes in marine sediments (Wuchter et al., 2006; Roussel et al., 2009; Durbin and Teske, 2010). Archaea have key roles in marine biogeochemical cycles, particularly the carbon and nitrogen cycles, and they also contribute to sulfur cycling in marine sediments (Auguet et al., 2010; Offre et al., 2013; Zhang et al., 2015; Danovaro et al., 2016; He et al., 2016).
Archaea were first reported in the cold ocean in 1992 (DeLong, 1992; Fuhrman et al., 1992), and this led to the identification of two major groups based on 16S rRNA sequences: Marine Group Ⅰ (MG-Ⅰ, now classified as Nitrosopumilales in Phylum Thaumarchaeota) and Marine Group Ⅱ (MG-Ⅱ, belonging to Phylum Euryarchaeota) (Brochier-Armanet et al., 2008; Zhang et al., 2015; Qin et al., 2017). The MG-Ⅰ are the most abundant and cosmopolitan chemoautotrophic archaea in the marine environment (Bano et al., 2004; Offre et al., 2013). Studies on this group have reported that they can fix inorganic carbon and oxidize ammonia aerobically (Stahl and De La Torre, 2012). Therefore, these species may be responsible for a significant fraction of chemoautotrophic production in the dark ocean and could play a vital role in marine biogeochemical cycles, particularly the carbon and nitrogen cycles, and might be responsible for a significant amount of the primary production in dark ocean ecosystems (Herndl et al., 2005; Tourna et al., 2011; Offre et al., 2013). There is evidence that marine MG-Ⅰ might account for more than 10% of the total prokaryotic production in the ocean (Herndl et al., 2005; Offre et al., 2013). Previous studies have defined different phylogenetic ecotypes of MG-Ⅰ (Massana et al., 2000; Takai et al., 2004; Durbin and Teske, 2012; Jorgensen et al., 2012; Lauer et al., 2016). The MG-Ⅰ.a phylotype is common in shallow seawater and the MG-Ⅰ.c phylotype is common in deep seawater (Massana et al., 2000; Takai et al., 2004). MG-Ⅱ are also abundant in the marine photic zone, which lives heterotrophically, and might play important roles in the long-term storage of dissolved organic carbon (Zhang et al., 2015). Besides, members of Miscellaneous Crenarchaeota Group (MCG) affiliated in phylum Bathyarchaeota were frequently detected in the marine subsurface sediments (Meng et al., 2014; He et al., 2016).
Seamounts, defined as subsurface topographic rises at least 1 km above the seafloor, usually form from volcanoes in the deep sea, and often have peaks that are hundreds to thousands of meters below the surface (Menard, 1964). Seamounts occur at various depths in all oceans worldwide and vary in form and size (Schlacher et al., 2010). Seamounts are also hosts to unique oceanic ecosystems, and they provide a variety of habitats and environmental conditions for benthic organisms (Clark et al., 2010). Seamounts affect underwater currents and create upwellings that attract plankton, corals, and fish (Danovaro et al., 2009; Clark et al., 2010; Rowden et al., 2010), and are believed to support high levels of biodiversity and endemic marine animals (Schlacher et al., 2010). The benthic communities of seamounts are distinct from those of continents and oceanic islands, in that they support greater biomass and have different community compositions (Rowden et al., 2010; Yesson et al., 2011).
Seamounts potentially provide many unique niches for microorganisms, but little is known of the composition of Archaea in seamounts area. There have been many studies of Archaea assemblages near hydrothermal vents (Huber et al., 2002; Nakagawa et al., 2004; Hara et al., 2005; Opatkiewicz et al., 2009; Kato et al., 2013) and ferromanganese crusts (Liao et al., 2011; Nitahara et al., 2011; Huo et al., 2015; Nitahara et al., 2017). Phylotypes in MG-Ⅰ were predominant, and those of MG-Ⅱ, MG-Ⅲ, and Marine Benthic Group E (MBGE) were also reported in the hydrothermal fluids and waters of Suiyo Seamount and hydrothermal vents at Axial Seamount, in the Juan de Fuca Ridge (Huber et al., 2002; Nakagawa et al., 2004; Hara et al., 2005; Opatkiewicz et al., 2009; Kato et al., 2013). Characterization of microbial populations in sediments of the Marsili and Palinuro Seamounts in the Tyrrhenian Sea (Western Mediterranean) indicated that Crenarchaeota (now the Thaumarchaeota) dominated archaeal communities in seamount and non-seamount sediments, with an abundance generally 3 times higher than that of Euryarchaeota (Danovaro et al., 2009). Phylogenetic analyses showed the predominance of MG-Ⅰ in the sediments of cobaltrich ferromanganese crusts of central Pacific seamounts (Liao et al., 2011; Huo et al., 2015), and in the sediment and crust of the Takuyo-Daigo Seamount and the Ryusei Seamount (Nitahara et al., 2011; Nitahara et al., 2017). The MG-Ⅰ archaea appear to have roles in sulphur and nitrogen cycling (Huo et al., 2015). In summary, there are no systematic studies on Archaea inhabiting in seamount area, and based on these limited studies, MG-Ⅰ Thaumarchaeota was always predominant, and MG-Ⅱ, Euryarchaeota, and Marine Benthic Group E (MBGE) were also reported in seamount area.
As a unique ecosystem, it is important to better understand seamount biodiversity, the effect of different environmental parameters on the archaeal community and their function in the ecosystems. To fully understand the archaeal community in seamount, we performed a systematic investigation of the archaeal communities on a seamount of the Mariana Volcanic Arc, near the Challenger Deep, in the tropical western Pacific Ocean. We examined the community structure and distribution of Archaea and their relationships with different environmental variables in an effort to better understand microbial communities and their ecological functions in seamount ecosystems.
2 MATERIAL AND METHOD 2.1 Site description and sample collectionSediments were collected from the Yap Seamount area in the Western Pacific (139°26′–139°43′E, 11°26′–11°41′N) during a cruise of the Chinese R/V Kexue from Feb. to Apr. 2016. Surface sediment samples were collected a custom-made steel alloy collector using the remotely operated vehicle (ROV) Faxian (Discovery in Chinese) at 14 stations and depths of 238–2 024 m while assuring its safe operation (Fig. 1) (Liu et al., 2017). A sediment sample of approximately 10 g was collected from each station, transferred to a sterile sealed bag, and immediately stored at -20℃ for analyses of the archaeal community.
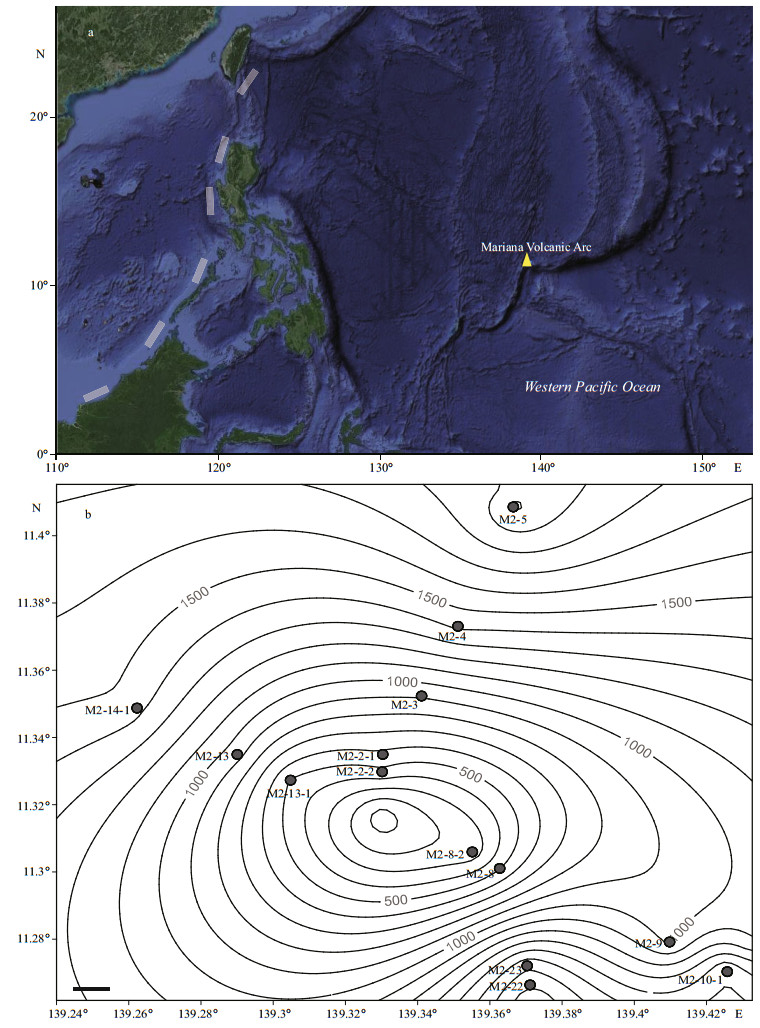
|
| Fig.1 Location and sampling stations of Kexue Seamount in the Mariana Volcanic Arc a. overview of the seamount location and Kexue Seamount was shown as a yellow triangle; b. 14 sampling stations were shown as black circles, and the scale bar is 1 km. |
Total genomic DNA was extracted directly from 0.5 g (wet weight) of mixed sediment using the FastDNA Spin Kit for Soil (MP Biomedicals, LLC, USA) and the Fast Prep FP120 Cell Disrupter (Thermo Electron Corp., USA). The quantity and quality of genomic DNA were determined using a NanoDrop ND-1000 spectrophotometer (LabTech, Washington, DC, USA) and 1.0% (w/v) agarose electrophoresis, respectively. The hypervariable region of the archaeal 16S rRNA gene was amplified using PCR (95℃ for 3 min; 35 cycles at 95℃ for 30 s, 55℃ for 30 s, and 72℃ for 45 s; a final extension at 72℃ for 10 min; and a cooling step at 10℃) with the bar-coded universal primers Arch344F (5′-barcode-ACGGGGYGCAGCAGGCGCGA-3′) and Arch915R (5′-barcode-GTGCTCCCCCGCCAATTCCT-3′) (Ohene-Adjei et al., 2007; Gantner et al., 2011). PCR was performed in triplicate in a 20- μL mixture, which included 4 μL of 5× Fast Pfu buffer, 2 μL of 2.5 mmol/L dNTPs, 0.8 μL of each primer (5 μmol/L), 0.4 μL of Fast Pfu polymerase, 0.2 μL of BSA, 10 ng of template DNA, and double distilled water (ddH2O). The PCR product was extracted from 2% agarose gels and purified using an AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA) according to the manufacturer's protocols.
2.3 Illumina MiSeq sequencing and operational taxonomic units assignmentEach purified PCR product was first quantified using a QuantiFluorTM-ST fluorometer with UV/Blue Channels (Promega, USA). Pair-end libraries were constructed using the TruSeq Nano DNA Library Prep Kit (Illumina, San Diego, CA, USA), and highthroughput sequencing was performed on the MiSeq (Illumina, San Diego, CA, USA) Sequencing Platform (Majorbio Bio-Pharm Technology Co. Ltd., Shanghai, China) following the manufacturer's protocols.
Raw FASTQ files were demultiplexed, qualityfiltered using QIIME (ver. 1.7.0 http://qiime.org/) (Caporaso et al., 2010) and Trimmomatic software (Bolger et al., 2014) with the following criteria: (ⅰ) 300 bp reads were truncated at any site receiving an average quality score below 20 over a 50-bp sliding window, and truncated reads shorter than 50 bp were discarded; (ⅱ) exact barcode matching was performed, with 2 nucleotide mismatch in primer matching, and reads containing ambiguous characters were removed; (ⅲ) only sequences that overlapped by more than 10 bp were assembled according to their overlap sequence. Reads that could not be assembled were discarded.
Operational taxonomic units (OTUs) were clustered using UPARSE (ver. 7.1 http://drive5.com/uparse/) (Edgar, 2013) with a 97% similarity cutoff, and chimeric sequences were identified and removed using UCHIME (http://drive5.com/uchime/) (Edgar et al., 2011) The classification of each OTU was determined by alignment with the SILVA database (Version 132 http://www.arb-silva.de), and using the RDP Classifier tool (ver. 2.2 http://sourceforge.net/projects/rdp-classifier/) with a credibility threshold of 70% (Wang et al., 2007; Quast et al., 2012). Taxonomic groups were derived from the number of reads assigned to each taxon at all ranks, from domain to genus, using QIIME. Bar graphs were plotted using an R software package, and a Circos graph (chord diagram) for archaeal community composition was created using Circos software (ver. 0.67 http://circos.ca/) (Krzywinski et al., 2009).
2.4 Diversity estimations, statistical and phylogenetic analysisTo equalize the sequencing effort, reads were randomly resampled using the number of reads obtained at the station with the fewest reads using a Perl script (Gilbert et al., 2009). Mothur (ver. 1.30.1, http://www.mothur.org) was applied to estimate Alpha diversity (Schloss et al., 2009). Chao 1 and the abundance-based coverage estimator (ACE) were used to calculate community richness (Chao, 1984; Chao and Lee, 1992; Chao et al., 1993) and the Shannon and Simpson diversity indexes were used to characterize community diversity (Magurran, 1988; Simpson et al., 2002). Good's coverage at the 97% similarity level was used to evaluate effect of sample depth. Rarefaction curves were drawn from the calculated results of Mothur and plot-rarefaction using an R software package (Amato et al., 2013).
Correlations between archaeal clades and environmental variables were calculated by Pearson correlation analysis (two-tail test) using SPSS software. For beta diversity, the Bray-Curtis dissimilarity was calculated by QIIME, and a hierarchical clustering tree was plotted using an R software package, which clustered samples with similar OTU compositions and abundances (Bray and Curtis, 1957; Yoshioka, 2008). Principal component analysis (PCA) was performed using the R software base package (Jolliffe, 2002). Detrended correspondence analysis (DCA) and redundancy analysis (RDA) were conducted using Canoco (ver. 4.5) to assess the relationship between microbial assemblages and environmental factors (Hill and Gauch, 1980; Šmilauer and Lepš, 2014; Desarbo et al., 2016).
The 16S rRNA gene sequences of the most common 50 OTUs were aligned using CLUSTAL W multiple alignment (Thompson et al., 1994). A phylogenetic tree was constructed using MEGA software (Tamura et al., 2013), based on the neighbor-joining (NJ) algorithm, and bootstrap values were calculated from 1 000 replicates. The sequences of MG- I were defined using different clusters, as previously described (Massana et al., 2000; Takai et al., 2004; Durbin and Teske, 2012; Jorgensen et al., 2012; Lauer et al., 2016).
3 RESULT 3.1 Environmental characteristicsAll sampling stations are located on the Kexue Seamount in the Mariana Volcanic Arc (Fig. 1, Table S1). The peak of the seamount is approximately 20 m below the water surface, and the depth of the sample sites ranged from 238 to 2 024 m. Stations M2-22 and M2-23 were located at the southeast steep slope and differ from other sites in that they have high concentrations of trace elements. The sediments mainly consisted of different proportions of coral and foraminifera sands. The temperature of sediments at the deeper stations (>1 000 m) was below 4℃, and the temperatures of other stations ranged from 4.9℃ to 7.3℃. There were no significant differences in the salinity of the core water, but it was slightly lower at the deeper stations. The level of dissolved oxygen (DO) ranged from 1.5 to 4.9 mg/L, and the highest value was at station M2-8 (Table S1). Pearson correlation analysis indicated that temperature and salinity had strong negative correlations with depth (P < 0.01) (Table S2).
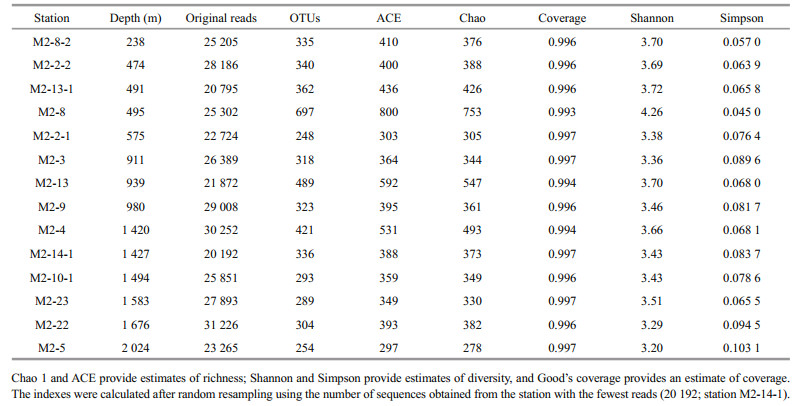
3.2 Archaeal diversity and richnessWe examined archaeal communities in the sediments from the different seamount sample stations by analysis of the 16S rRNA gene using amplicon sequencing. After quality control, we obtained 358 160 valid reads (average length: 532 bp; range: 20 192 to 31 226 reads per station) (Table 1). To equalize sequencing efforts, reads were randomly resampled using the number of reads obtained at the station with the fewest reads (20 192; station M2-14- 1). A total of 1 411 OTUs (based on 97% sequence similarity) were clustered after random resampling, and there were 248 to 697 OTUs per station (Table 1). The rarefaction curves indicated that sufficient sequencing intensities were achieved and that the populations were well-represented (Fig.S1). Good's coverage estimates for all samples exceeded 0.99 (Table 1), indicating that most species in the study area were represented by the amplicon sequencing provided by Illumina MiSeq sequencing. In general, there were decreases in community diversity (based on Shannon and Simpson indexes) and in species richness (based on Chao and ACE indexes) with increasing depth. In particular, the greatest diversity and richness were at station M2-8 (495 m), near the top of the seamount; the lowest diversity and richness were at station M2-5 (2 024 m), the deepest station (Table 1). Pearson correlation analysis indicated a negative correlation between the Shannon index and depth (P < 0.05), and a positive correlation between the Simpson index and depth (P < 0.05) (Table S2). In sum, the archaeal community diversity and species richness had a significant relationship with depth.
|
We performed taxonomic assignment at various levels using the Silva database. At the phylum level, the valid reads were mainly affiliated in the Thaumarchaeota (96.5%) and Nanoarchaeaeota (1.4%), and there were a small number of Euryarchaeota (0.2%), Crenarchaeota, Diapherotrites and Altiarchaeota. Thaumarchaeota was the most abundant phylum at all sample stations, and accounted for 87.2% (M2-8; 495 m) to 99.7% (M2-5; 2 024 m) of all reads, with an average of 96.5% (Fig. 2). Nanoarchaeaeota was the second most abundant phylum at all stations, and this phylum had the greatest abundance at stations M2-8 (5.3%), M2-13-1 (4.7%), M2-13 (3.0%), and M2-2-2 (2.3%), and M2-8-2 (1.0%), all of which were located near the top of the seamount (Fig. 2). The abundance of this phylum was less than 1% at all other stations. Euryarchaeota accounted for less than 1% of community abundance at all 14 stations.

|
| Fig.2 Archaeal community composition in sediments of the 14 seamount sample sites at the level of Phylum, Class, Order, and Genus, the average abundance is shown in parentheses |
At the class taxonomic level, Nitrososphaeria (belonging to Thaumarchaeota) dominated at all the 14 stations, and ranged from 83.1% (M2-8) to 99.0% (M2-2-1), with an average of 96.5%. Marine Benthic Group A (MBGA) had the highest abundance of 6.1% at station M2-13 and had higher abundances at stations M2-2-2, M2-13-1, and M2-4 (Fig. 2; yellow bars). Woesearchaeia had an abundance of 5.3% at station M2-8, and higher abundance at M2-13-1, M2- 13, M2-2-2, which were located near the top of the seamount (Fig. 2; violet bars). Additional lineages, such as Group 1.1c belonging to phylum Thaumarchaeota, Nanohaloarchaeia belonging to phylum Nanoarchaeaeota, Thermoplasmata belonging to phylum Euryarchaeota, had abundances less than 1% at all stations (Fig. 2).
At an order level, Nitrosopumilales (originally termed MG-Ⅰ) dominated at all 14 stations and ranged from 82.4% (M2-8) to 99.0% (M2-2-1). Except for unclassified reads, reads in were Nitrosopumilales grouped into three genera, Candidatus Nitrosopumilus, Candidatus Nitrosopelagicus and Cenarchaeum belonging to family Nitrosopumilaceae. Unclassified reads were dominant at all stations except M2-22 (Fig. 2; red bars); while Candidatus Nitrosopumilus accounted for 50.7% and unclassified reads accounted for 47.2% at this station. Notably, the abundance of Candidatus Nitrosopumilus tended to increase with depth and had the greatest abundance at stations M2- 22, M2-23, M2-10-1, M2-5, and M2-14-1 (Fig. 2; green bars). At station M2-8-2, Cenarchaeum accounted for approximately 1.2% of community abundance, which was 2- to 100-fold higher than at other stations (Fig. 2; dark red bars). Notably, there were only several MG-Ⅱ reads were found in the seamount, with an average of 0.05 %. The highest abundance (0.23%) occurred at a shallower station (M2-8, 495 m).
A Circos graph of community composition shows that unclassified Nitrosopumilales dominated the archaeal community at all stations except M2-22 where Nitrosopumilus had a high abundance (Fig. 3). Community composition analysis indicated that stations at different depths and locations had different community structures. In particular, Candidatus Nitrosopumilus was dominant at greater depth stations with lower temperature, but unclassified Nitrosopumilales, MBGA, dominated near the top of the seamount. There was a unique archaeal composition at the station on the steep southeast slope relative to most other stations.

|
| Fig.3 Circos graph (chord diagram) of archaeal community composition, showing the relationships of the main archaeal groups at different sample stations |
To simplify the analysis, we excluded rare organisms and analyzed the most abundant 50 OTUs (read numbers of 736 to 54 358), which accounted for more than 95% of total reads. Among these 50 OTUs, Nitrosopumilales accounted for 48 OTUs, and Marine Benthic Group A (MBGA) accounted for 2 OTUs. Phylogenetic analysis of these 50 OTUs showed that MG-Ⅰ was in 7 known clusters, and mainly affiliated with ε-ζ-θ, α, and υ clusters. There were also 6 OTUs in 4 unknown clusters, indicating the possible presence of unique archaeal species in the seamount (Fig. 4).
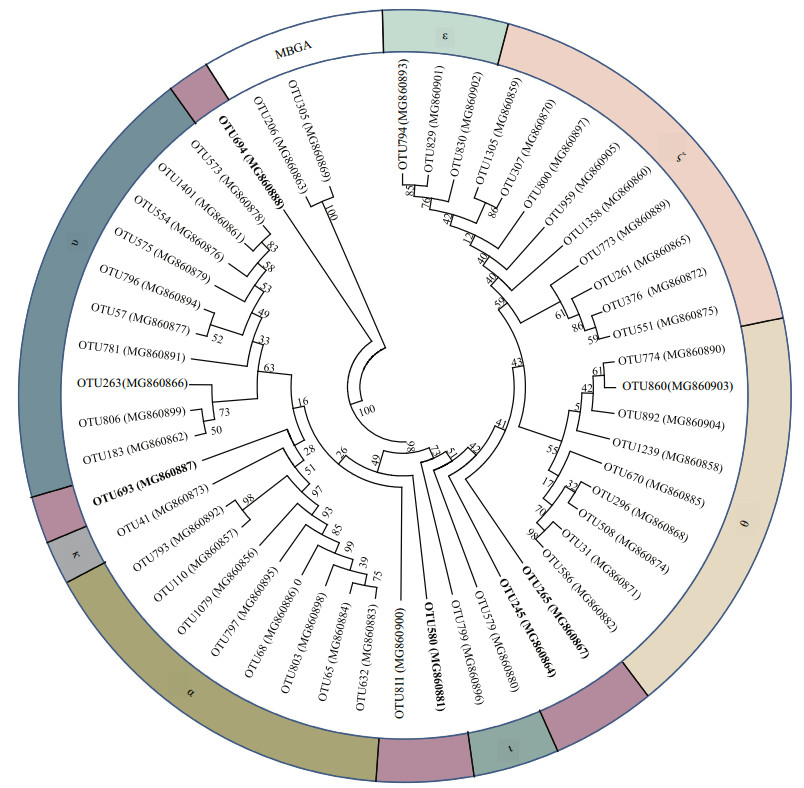
|
| Fig.4 Phylogenetic relationships of the most abundant 50 archaeal OTUs (~450 bp for alignment length) in the seamount based on 16S rRNA gene sequences, using two MBGA OTUs as out-groups The tree was constructed using the neighbor-joining (NJ) algorithm, and bootstrap values (from 1 000 replicates) are shown as percentages at branch points. GenBank accession numbers of the sequences used are indicated in parentheses. |
Analysis of beta diversity based on OTU composition and abundance showed that all but two of the seamount stations (M2-8-2 at 238 m and M2-22 at 1 676 m) grouped the archaeal communities into three clusters (Fig. 5). One cluster had stations with water depths less than 500 m (M2-2-1, M2-8, M2-13- 1, and M2-2-2), a second cluster had stations at depths of 500–1 000 m (M2-3, M2-13, and M2-9; M2-4 with a depth of 1 420 m), and a third cluster had 4 stations at depths greater than 1 000 m (M2-23, M2-10-1, M2- 5, and M2-14-1). Principal component analysis (PCA) indicated a similar grouping of sample stations (Fig. 6). These results thus showed a clear transition of archaeal community composition as a function of water depth in this seamount.
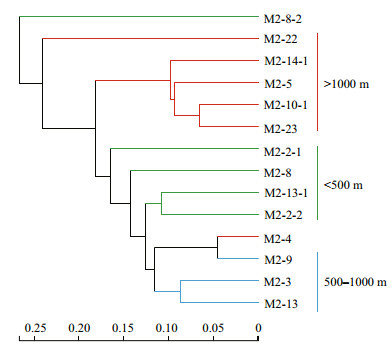
|
| Fig.5 Bray-Curtis hierarchical cluster tree of OTU composition and abundance of Archaea in the seamount Green: water depth < 500 m; blue: water depth=500-1 000 m; red: water depth >1 000 m. |

|
| Fig.6 Principal component analysis of archaeal communities based on OTU composition and abundance |
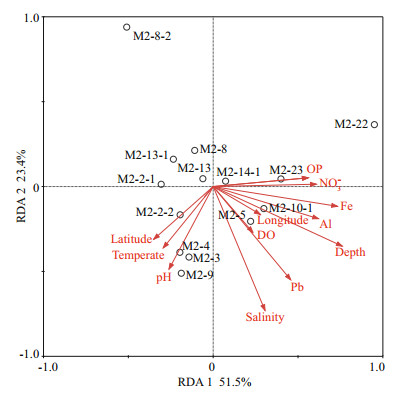
We also performed a detrended correspondence analysis (DCA) for all samples to examine the effect of environmental variables on community composition. The longest gradient length was 2.078, indicating that all samples were suitable for redundancy analysis (RDA) using a linear model. The different environmental parameters were selected using a Monte-Carlo test, and a P-value below 0.05 was used as the standard. The first and second RDA components explained 74.9% of the variability in archaeal community structure. Moreover, community divergence mainly correlated with depth, salinity, and concentration of Pb. Dissolved oxygen (DO) and latitude gradients had no clear effects on community structure (Fig. 7).
|
| Fig.7 Detrended correspondence analysis of the relationship between archaeal community structure at the OTU level according to sample site and environmental variables Environmental variables are indicated by arrows, and sample sites as open circles. |
We then conducted correlation analysis to assess the relationship of different environmental variables with Archaea families and individual OTUs. Pearson analysis indicated that the abundance of Thaumarchaeota correlated positively with water depth and Pb concentration (P < 0.05) and that the abundance of Woesearchaeota correlated negatively with water depth and Pb concentration (P < 0.05). Pearson analysis also showed that abundance of Nitrosopumilus correlated positively with Fe concentration (P < 0.01), depth (P < 0.05), and Al concentration (P < 0.05) (Table S2).
4 DISCUSSIONArchaea are now recognized as important as bacteria in marine biogeochemical cycling. Archaea are not only among the most abundant microbial plankton groups in the oceans but are also highly abundant in marine sediments (DeLong, 1992; Bano et al., 2004), (Wuchter et al., 2006; Roussel et al., 2009; Durbin and Teske, 2010).
In the present study, we investigated the archaeal diversity and communities of seamount in the Mariana Volcanic Arc, the western Pacific Ocean using highthroughput sequencing. Interestingly, we found that community diversity and species richness decreased with depth, with the highest values at station M2-8 (495 m), near the top of the seamount, and the lowest values at station M2-5 (2 024 m), the deepest sample station. A clear transition of archaeal community composition as a function of water depth could be found in this seamount, so this distinct community composition might be generally consistent with depth. The depth-dependent changes in the distribution of Archaea that we observed might be due to the temperature, which also decreased with depth. Seamounts rise abruptly from the ocean seafloor and have a substantial impact on physical oceanography. In particular, they affect the nature of circulation, local hydrographic distributions, turbulence, and vertical and horizontal transport of nutrients (Lavelle and Mohn, 2010). It seems reasonable that organic matter flux could have an influence on the distribution, as the observed decreased temperature, having an effect on the energy flux and activity. The seamount flows also have an important influence on bacterial community structure were reported (Emerson and Moyer, 2010), as does depth (O'Hara, 2007; Clark et al., 2010). So, the temperature and seamount circulation might jointly lead to the depth-dependent diversity and richness changes in the distribution of Archaea.
MG-Ⅰ (now classified as Nitrosopumilales in phylum Thaumarchaeota) and MG-Ⅱ (belonging to phylum Euryarchaeota) are the two major phyla of marine Archaea based on analysis of 16S rRNA sequences. In our study, we found that Nitrosopumilales was dominant in surface sediment, most in α, κ-υ, and ε-ζ-θ clusters. Our analysis of a subgroup of Nitrosopumilales sequences indicated the presence of 4 unknown clusters, suggesting there might be novel archaeal species in this seamount. There were still a small amount of Woesearchaeia and Marine Benthic Group A archaea reads in Kexue Seamount. However, there were only several MG-Ⅱ reads were found, with an average of 0.05%. The highest abundance (0.23%) occurred at a shallower station (M2-8, 495 m).
Previous studies of different seamounts had reported that phylotypes affiliated within MG-Ⅰ were always dominant, and most individuals are in the α, η-κ-υ, and ε-ζ-θ clusters (Nakagawa et al., 2004; Hara et al., 2005; Kato et al., 2013). Ammonia-oxidizing Archaea (AOA) in MG-Ⅰ, which have the amoA gene, produce energy from the aerobic oxidation of ammonia (NH4+) to nitrite (NO2ˉ) (Stahl and De La Torre, 2012). CO2 is the main carbon source of MG-Ⅰ, but co-assimilation of organic carbon can also occur (Tourna et al., 2011). This means that MG-Ⅰ organisms play a vital role in marine biogeochemical cycles, particularly the carbon and nitrogen cycles, and might be responsible for a significant amount of the primary production in dark ocean ecosystems (Herndl et al., 2005; Offre et al., 2013). MG-Ⅰ have a global distribution and are abundant in marine water and sediments (Brochier-Armanet et al., 2008, Stahl and De La Torre, 2012). MG-Ⅱ and other Euryarchaeota archaea were also present in seamount, with the highest abundance of 30% (Nakagawa et al., 2004; Kato et al., 2013), whilst, there were a very small amount of MG-Ⅱ were found in our study. MG-Ⅱ is abundant in marine aquatic environments and mainly occur in the photic zone. Their diversity is also greatest in surface seawater and decreases with depth. These organisms live heterotrophically, and are important in the long-term storage of dissolved organic carbon (Zhang et al., 2015).
Seamounts provide a variety of habitats and environmental conditions for benthic organisms (Clark et al., 2010), and affect underwater currents and create upwellings that attract plankton, corals, and fish (Danovaro et al., 2009; Clark et al., 2010; Rowden et al., 2010). Seamount is believed to support high levels of biodiversity and endemic marine animals (Schlacher et al., 2010). Biomass deposition from topwater and a large diversity of organism would promote the accumulation of organic matter in seamount ecosystem. MG-Ⅰ archaea can drive the assimilation of organic carbon, but no evidence indirectly degrade organic matter. Meanwhile, MG-Ⅱ archaea unlikely to participate in organic matter degradation, because of their absence in sediment. We hypothesize that MG-Ⅰ archaea might play an indirect role in organic matter degradation progress, with assisting bacteria in top sediment. Which group of the microbiological organism could contribute to the degradation of organic matter needs to be further study, comprehensively considering the contribution of all the trophic levels in microbial net.
Moreover, the site of the present study is a seamount at the junction of the Yap and Mariana trenches, In particular, the seamount sediments mainly consist of coral and foraminifera sands, which have lower densities of organic carbon (Liu et al., 2017). Because of the different requirements of MG-Ⅰ and MG-Ⅱ, the unique distribution patterns of Archaea in this seamount might be a consequence of the sediment heterogeneity and the complex geochemistry. The geographic properties of the seamount might contribute to the unique community structure, and made it different from microbial communities of other seamounts.
5 CONCLUSIONCompared with other ocean ecosystems, there is only limited information of Archaea in seamount habitats, which in general are not well-studied environments. We reported the first systematic analysis of archaeal diversity and community structure in a seamount habitat by use of high-throughput sequencing. The results indicated that Nitrosopumilales was the dominant clade at all 14 sample stations, and that water depth and geographic variables seemed to be important in shaping the distinctive archaeal communities. Our study provides a foundation for the study of Archaea in seamounts, and the unique distribution patterns also shed a light on the ecological functions of Archaea in seamounts.
6 DATA AVAILABILITY STATEMENTThe 16S rRNA gene sequences of the most abundant 50 OTUs in the current study were deposited in the NCBI under accession numbers MG860856- MG860905.
7 ACKNOWLEDGMENTWe thank the crew of the R/V Kexue, the ROV team, and W. Zhang for collecting sediment samples.
Electronic supplementary materialSupplementary material (Supplementary Tables S1–S2, Fig.S1) is available in the online version of this article at https://doi.org/10.1007/s00343-019-8044-x.
Amato K R, Yeoman C J, Kent A, Righini N, Carbonero F, Estrada A, Gaskins H R, Stumpf R M, Yildirim S, Torralba M, Gillis M, Wilson B A, Nelson K E, White B A, Leigh S R. 2013. Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. The ISME Journal, 7(7): 1344-1353.
DOI:10.1038/ismej.2013.16 |
Auguet J C, Barberan A, Casamayor E O. 2010. Global ecological patterns in uncultured Archaea. The ISME Journal, 4(2): 182-190.
DOI:10.1038/ismej.2009.109 |
Bano N, Ruffin S, Ransom B, Hollibaugh J T. 2004. Phylogenetic composition of Arctic Ocean archaeal assemblages and comparison with Antarctic assemblages. Applied and Environmental Microbiology, 70(2): 781-789.
DOI:10.1128/aem.70.2.781-789.2004 |
Bolger A M, Lohse M, Usadel B. 2014. Trimmomatic:a flexible trimmer for Illumina sequence data. Bioinformatics, 30(15): 2114-2120.
DOI:10.1093/bioinformatics/btu170 |
Bray J R, Curtis J T. 1957. An ordination of the upland forest communities of southern Wisconsin. Ecological Society of America, 27(4): 325-349.
DOI:10.2307/1942268 |
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P. 2008. Mesophilic Crenarchaeota:proposal for a third archaeal phylum, the Thaumarchaeota. Nature Reviews Microbiology, 6(3): 245-252.
DOI:10.1038/nrmicro1852 |
Caporaso J G, Kuczynski J, Stombaugh J, Bittinger K, Bushman F D, Costello E K, Fierer N, Peña A G, Goodrich J K, Gordon J I, Huttley G A, Kelley S T, Knights D, Koenig J E, Ley R E, Lozupone C A, McDonald D, Muegge B D, Pirrung M, Reeder J, Sevinsky J R, Turnbaugh P J, Walters W A, Widmann J, Yatsunenko T, Zaneveld J, Knight R. 2010. QⅡME allows analysis of high-throughput community sequencing data. Nature Methods, 7(5): 335-336.
DOI:10.1038/nmeth.f.303 |
Chao A, Lee S M. 1992. Estimating the number of classes via sample coverage. Journal of the American Statistical Association, 87(417): 210-217.
DOI:10.1080/01621459.1992.10475194 |
Chao A, Ma M C, Yang M C K. 1993. Stopping rules and estimation for recapture debugging with unequal failure rates. Biometrika, 80(1): 193-201.
DOI:10.1093/biomet/80.1.193 |
Chao A. 1984. Nonparametric estimation of the number of classes in a population. Scandinavian Journal of Statistics, 11(4): 265-270.
|
Clark M R, Rowden A A, Schlacher T, Williams A, Consalvey M, Stocks K I, Rogers A D, O'Hara T D, White M, Shank T M, Hall-Spencer J M. 2010. The ecology of seamounts:structure, function, and human impacts. Annual Review of Marine Science, 2: 253-278.
DOI:10.1146/annurev-marine-120308-081109 |
Danovaro R, Corinaldesi C, Luna G M, Magagnini M, Manini E, Pusceddu A. 2009. Prokaryote diversity and viral production in deep-sea sediments and seamounts. DeepSea Research Part Ⅱ:Topical Studies in Oceanography, 56(11-12): 738-747.
DOI:10.1016/j.dsr2.2008.10.011 |
Danovaro R, Molari M, Corinaldesi C, Dell'Anno A. 2016. Macroecological drivers of archaea and bacteria in benthic deep-sea ecosystems. Science Advances, 2(4): e1500961.
DOI:10.1126/sciadv.1500961 |
DeLong E F. 1992. Archaea in coastal marine environments. Proceedings of the National Academy of Sciences of the United States of America, 89(12): 5685-5689.
DOI:10.1073/pnas.89.12.5685 |
DeSarbo W S, Hwang H, Jedidi K. 2016. Redundancy Analysis. John Wiley & Sons, Ltd..
|
Durbin A M, Teske A. 2010. Sediment-associated microdiversity within the Marine Group Ⅰ Crenarchaeota. Environmental Microbiology Reports, 2(5): 693-703.
DOI:10.1111/j.1758-2229.2010.00163.x |
Durbin A M, Teske A. 2012. Archaea in organic-lean and organic-rich marine subsurface sediments:an environmental gradient reflected in distinct phylogenetic lineages. Frontiers in Microbiology, 3: 168.
DOI:10.3389/fmicb.2012.00168 |
Edgar R C, Haas B J, Clemente J C, Quince C, Knight R. 2011. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics, 27(16): 2194-2200.
DOI:10.1093/bioinformatics/btr381 |
Edgar R C. 2013. UPARSE:highly accurate OTU sequences from microbial amplicon reads. Nature Methods, 10(10): 996-998.
DOI:10.1038/nmeth.2604 |
Emerson D, Moyer C L. 2010. Microbiology of seamounts:common patterns observed in community structure. Oceanography, 23(1): 148-163.
DOI:10.5670/oceanog.2010.67 |
Fuhrman J A, McCallum K, Davis A A. 1992. Novel major archaebacterial group from marine plankton. Nature, 356(6365): 148-149.
DOI:10.1038/356148a0 |
Gantner S, Andersson A F, Alonso-Sáez L, Bertilsson S. 2011. Novel primers for 16S rRNA-based archaeal community analyses in environmental samples. Journal of Microbiological Methods, 84(1): 12-18.
DOI:10.1016/j.mimet.2010.10.001 |
Gilbert J A, Field D, Swift P, Newbold L, Oliver A, Smyth T, Somerfield P J, Huse S, Joint I. 2009. The seasonal structure of microbial communities in the Western English Channel. Environmental Microbiology, 11(12): 3132-3139.
DOI:10.1111/j.1462-2920.2009.02017.x |
Hara K, Kakegawa T, Yamashiro K, Maruyama A, Ishibashi J I, Marumo K, Urabe T, Yamagishi A. 2005. Analysis of the archaeal sub-seafloor community at Suiyo Seamount on the Izu-Bonin Arc. Advances in Space Research, 35(9): 1634-1642.
DOI:10.1016/j.asr.2005.04.111 |
He Y, Li M, Perumal V, Feng X, Fang J, Xie J, Sievert S M, Wang F. 2016. Genomic and enzymatic evidence for acetogenesis among multiple lineages of the archaeal phylum Bathyarchaeota widespread in marine sediments. Nature Microbiology, 1(6): 16035.
DOI:10.1038/nmicrobiol.2016.35 |
Herndl G J, Reinthaler T, Teira E, Van Aken H, Veth C, Pernthaler A, Pernthaler J. 2005. Contribution of Archaea to total prokaryotic production in the deep Atlantic Ocean. Applied and Environmental Microbiology, 71(5): 2303-2309.
DOI:10.1128/AEM.71.5.2303-2309.2005 |
Hill M O, Gauch Jr H G. 1980. Detrended correspondence analysis:an improved ordination technique. Vegetatio, 42(1-3): 47-58.
DOI:10.1007/bf00048870 |
Huber J A, Butterfield D A, Baross J A. 2002. Temporal changes in archaeal diversity and chemistry in a midocean ridge subseafloor habitat. Applied and Environmental Microbiology, 68(4): 1585-1594.
DOI:10.1128/aem.68.4.1585-1594.2002 |
Huo Y Y, Cheng H, Post A F, Wang C S, Jiang X W, Pan J, Wu M, Xu X W. 2015. Ecological functions of uncultured microorganisms in the cobalt-rich ferromanganese crust of a seamount in the central Pacific are elucidated by fosmid sequencing. Acta Oceanologica Sinica, 34(4): 92-113.
DOI:10.1007/s13131-015-0650-7 |
Jolliffe I T. 2002. Principal Component Analysis. 2nd edn. Springer-Verlag, New York.
|
Jorgensen S L, Hannisdal B, Lanzén A, Baumberger T, Flesland K, Fonseca R, Øvreås L, Steen I H, Thorseth I H, Pedersen R B, Schleper C. 2012. Correlating microbial community profiles with geochemical data in highly stratified sediments from the Arctic Mid-Ocean Ridge. Proceedings of the National Academy of Sciences of the United States of America, 109(42): E2846-E2855.
DOI:10.1073/pnas.1207574109 |
Karner M B, DeLong E F, Karl D M. 2001. Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature, 409(6819): 507-510.
DOI:10.1038/35054051 |
Kato S, Nakawake M, Kita J, Yamanaka T, Utsumi M, Okamura K, Ishibashi J I, Ohkuma M, Yamagishi A. 2013. Characteristics of microbial communities in crustal fluids in a deep-sea hydrothermal field of the Suiyo Seamount. Frontiers in Microbiology, 4: 85.
DOI:10.3389/fmicb.2013.00085 |
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones S J, Marra M A. 2009. Circos:an information aesthetic for comparative genomics. Genome Research, 19(9): 1639-1645.
DOI:10.1101/gr.092759.109 |
Lauer A, Sorensen K B, Teske A. 2016. Phylogenetic characterization of marine benthic Archaea in organicpoor sediments of the eastern Equatorial Pacific Ocean(ODP Site 1225). Microorganisms, 4(3): 32.
DOI:10.3390/microorganisms4030032 |
Lavelle J W, Mohn C. 2010. Motion, commotion, and biophysical connections at deep ocean seamounts. Oceanography, 23(1): 90-103.
DOI:10.5670/oceanog.2010.64 |
Liao L, Xu X W, Jiang X W, Wang C S, Zhang D S, Ni J Y, Wu M. 2011. Microbial diversity in deep-sea sediment from the cobalt-rich crust deposit region in the Pacific Ocean. FEMS Microbiology Ecology, 78(3): 565-585.
DOI:10.1111/j.1574-6941.2011.01186.x |
Liu J, Zhang W Y, Li X G, Li X G, Chen X M, Li J H, Teng Z J, Xu C, Santini C L, Zhao L, Zhao Y, Zhang H, Zhang W J, Xu K D, Li C L, Pan Y X, Xiao T, Pan H M, Wu L F. 2017. Bacterial community structure and novel species of magnetotactic bacteria in sediments from a seamount in the Mariana volcanic arc. Scientific Reports, 7(1): 17964.
DOI:10.1038/s41598-017-17445-4 |
Magurran A E. 1988. Ecological Diversity and Its Measurement. Princeton University Press, Princeton, USA.
|
Massana R, DeLong E F, Pedrós-Alió C. 2000. A few cosmopolitan phylotypes dominate planktonic archaeal assemblages in widely different oceanic provinces. Applied and Environmental Microbiology, 66(5): 1777-1787.
DOI:10.1128/AEM.66.5.1777-1787.2000 |
Menard H W. 1964. Marine Geology of the Pacific. McGrawHill, New York, NY.
|
Meng J, Xu J, Qin D, He Y, Xiao X, Wang F P. 2014. Genetic and functional properties of uncultivated MCG archaea assessed by metagenome and gene expression analyses. The ISME Journal, 8(3): 650-659.
DOI:10.1038/ismej.2013.174 |
Nakagawa T, Ishibashi J I, Maruyama A, Yamanaka T, Morimoto Y, Kimura H, Urabe T, Fukui M. 2004. Analysis of dissimilatory sulfite reductase and 16S rRNA gene fragments from deep-sea hydrothermal sites of the Suiyo Seamount, Izu-Bonin Arc, Western Pacific. Applied and Environmental Microbiology, 70(1): 393-403.
DOI:10.1128/AEM.70.1.393-403.2004 |
Nitahara S, Kato S, Urabe T, Usui A, Yamagishi A. 2011. Molecular characterization of the microbial community in hydrogenetic ferromanganese crusts of the Takuyo-Daigo Seamount, northwest Pacific. FEMS Microbiology Letters, 321(2): 121-129.
DOI:10.1111/j.1574-6968.2011.02323.x |
Nitahara S, Kato S, Usui A, Urabe T, Suzuki K, Yamagishi A. 2017. Archaeal and bacterial communities in deep-sea hydrogenetic ferromanganese crusts on old seamounts of the northwestern Pacific. PLoS One, 12(2): e0173071.
DOI:10.1371/journal.pone.0173071 |
O'Hara T D. 2007. Seamounts:centres of endemism or species richness for ophiuroids?. Global Ecology and Biogeography, 16(6): 720-732.
DOI:10.1111/j.1466-8238.2007.00329.x |
Offre P, Spang A, Schleper C. 2013. Archaea in biogeochemical cycles. Annual Review of Microbiology, 67: 437-457.
DOI:10.1146/annurev-micro-092412-155614 |
Ohene-Adjei S, Teather R M, Ivan M, Forster R J. 2007. Postinoculation protozoan establishment and association patterns of methanogenic archaea in the ovine rumen. Applied and Environmental Microbiology, 73(14): 4609-4618.
DOI:10.1128/AEM.02687-06 |
Opatkiewicz A D, Butterfield D A, Baross J A. 2009. Individual hydrothermal vents at Axial Seamount harbor distinct subseafloor microbial communities. FEMS Microbiology Ecology, 70(3): 413-424.
DOI:10.1111/j.1574-6941.2009.00747.x |
Qin W, Heal K R, Ramdasi R, Kobelt J N, Martens-Habbena W, Bertagnolli A D, Amin S A, Walker C B, Urakawa H, Könneke M, Devol A H, Moffett J W, Armbrust E V, Jensen G J, Ingalls A E, Stahl D A. 2017. Nitrosopumilus maritimus gen. nov., sp. nov., Nitrosopumilus cobalaminigenes sp. nov., Nitrosopumilus oxyclinae sp.nov., and Nitrosopumilus ureiphilus sp. nov., four marine ammonia-oxidizing archaea of the phylum Thaumarchaeota. International Journal of Systematic and Evolutionary Microbiology, 67(12): 5067-5079.
DOI:10.1099/ijsem.0.002416 |
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner F O. 2012. The SILVA ribosomal RNA gene database project:improved data processing and web-based tools. Nucleic Acids Research, 41(D1): D590-D596.
DOI:10.1093/nar/gks1219 |
Roussel E G, Sauvadet A L, Chaduteau C, Fouquet Y, Charlou J L, Prieur D, Cambon Bonavita M A. 2009. Archaeal communities associated with shallow to deep subseafloor sediments of the New Caledonia Basin. Environmental Microbiology, 11(9): 2446-2462.
DOI:10.1111/j.1462-2920.2009.01976.x |
Rowden A A, Schlacher T A, Williams A, Clark M R, Stewart R, Althaus F, Bowden D A, Consalvey M, Robinson W, Dowdney J. 2010. A test of the seamount oasis hypothesis:seamounts support higher epibenthic megafaunal biomass than adjacent slopes. Marine Ecology, 31(S1): 95-106.
DOI:10.1111/j.1439-0485.2010.00369.x |
Schlacher T A, Rowden A A, Dower J F, Consalvey M. 2010. Seamount science scales undersea mountains:new research and outlook. Marine Ecology, 31(S1): 1-13.
DOI:10.1111/j.1439-0485.2010.00396.x |
Schloss P D, Westcott S L, Ryabin T, Hall J R, Hartmann M, Hollister E B, Lesniewski R A, Oakley B B, Parks D H, Robinson C J, Sahl J W, Stres B, Thallinger G G, Van Horn D J, Weber C F. 2009. Introducing mothur:opensource, platform-independent, community-supported software for describing and comparing microbial communities. Applied and Environmental Microbiology, 75(23): 7537-7541.
DOI:10.1128/AEM.01541-09 |
Simpson J M, Kocherginskaya S A, Aminov R I, Skerlos L T, Bradley T M, Mackie R I, White B A. 2002. Comparative microbial diversity in the gastrointestinal tracts of food animal species. Integrative and Comparative Biology, 42(2): 327-331.
DOI:10.1093/icb/42.2.327 |
Šmilauer P, Lepš J. 2014. Multivariate Analysis of Ecological Data Using Canoco 5. Cambridge University Press, Cambridge.
|
Stahl D A, De La Torre J R. 2012. Physiology and diversity of ammonia-oxidizing archaea. Annual Review of Microbiology, 66: 83-101.
DOI:10.1146/annurev-micro-092611-150128 |
Takai K, Oida H, Suzuki Y, Hirayama H, Nakagawa S, Nunoura T, Inagaki F, Nealson K H, Horikoshi K. 2004. Spatial distribution of marine Crenarchaeota group Ⅰ in the vicinity of deep-sea hydrothermal systems. Applied and Environmental Microbiology, 70(4): 2404-2413.
DOI:10.1128/aem.70.4.2404-2413.2004 |
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. 2013. MEGA6:molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30(12): 2725-2729.
DOI:10.1093/molbev/mst197 |
Thompson J D, Higgins D G, Gibson T J. 1994. CLUSTAL W:improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22(22): 4673-4680.
DOI:10.1093/nar/22.22.4673 |
Tourna M, Stieglmeier M, Spang A, Konneke M, Schintlmeister A, Urich T, Engel M, Schloter M, Wagner M, Richter A, Schleper C. 2011. Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proceedings of the National Academy of Sciences of the United States of America, 108(20): 8420-8425.
DOI:10.1073/pnas.1013488108 |
Wang Q, Garrity G M, Tiedje J M, Cole J R. 2007. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Applied and Environmental Microbiology, 73(16): 5261-5267.
DOI:10.1128/AEM.00062-07 |
Woese C R, Fox G E. 1977. Phylogenetic structure of the prokaryotic domain:the primary kingdoms. Proceedings of the National Academy of Sciences of the United States of America, 74(11): 5088-5090.
DOI:10.1073/pnas.74.11.5088 |
Woese C R, Kandler O, Wheelis M L. 1990. Towards a natural system of organisms:proposal for the domains Archaea, Bacteria, and Eucarya. Proceedings of the National Academy of Sciences of the United States of America, 87(12): 4576-4579.
DOI:10.1073/pnas.87.12.4576 |
Wuchter C, Abbas B, Coolen M J L, Herfort L, Van Bleijswijk J, Timmers P, Strous M, Teira E, Herndl G J, Middelburg J J, Schouten S, Sinninghe Damsté J S. 2006. Archaeal nitrification in the ocean. Proceedings of the National Academy of Sciences of the United States of America, 103(33): 12317-12322.
DOI:10.1073/pnas.0600756103 |
Yesson C, Clark M R, Taylor M L, Rogers A D. 2011. The global distribution of seamounts based on 30 arc seconds bathymetry data. Deep Sea Research Part Ⅰ:Oceanographic Research Papers, 58(4): 442-453.
DOI:10.1016/j.dsr.2011.02.004 |
Yoshioka P M. 2008. Misidentification of the Bray-Curtis similarity index. Marine Ecology Progress Series, 368: 309-310.
DOI:10.3354/meps07728 |
Zhang C L, Xie W, Martin-Cuadrado A B, Rodriguez-Valera F. 2015. Marine group Ⅱ Archaea, potentially important players in the global ocean carbon cycle. Frontiers in Microbiology, 6: 1108.
DOI:10.3389/fmicb.2015.01108 |
 2019, Vol. 37
2019, Vol. 37


