Institute of Oceanology, Chinese Academy of Sciences
Article Information
- ZANG Zhengrong, LI Yanhua, HU Qiang, HAN Danxiang
- Unraveling enhanced membrane lipid biosynthesis in Chlamydomonas reinhardtii starchless mutant sta6 by using an electrospray ionization mass spectrometry-based lipidomics method
- Journal of Oceanology and Limnology, 38(3): 783-794
- http://dx.doi.org/10.1007/s00343-019-9138-1
Article History
- Received May. 17, 2019
- accepted in principle Jul. 2, 2019
- accepted for publication Aug. 22, 2019
2 Key Laboratory for Algal Biology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China;
3 University of Chinese Academy of Sciences, Beijing 100049, China;
4 State Key Laboratory of Freshwater Ecology and Biotechnology, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan 430072, China;
5 Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory for Marine Science and Technology, Qingdao 266071, China;
6 Beijing Key Laboratory of Algae Biomass, Microalgae Biotechnology Center, SDIC Biotech Investment Co. Ltd., State Development & Investment Corp., Beijing 100142, China
The unicellular green microalga Chlamydomonas reinhardtii has been widely used as a model organism for studying lipid metabolism in photosynthetic organisms (Harris, 2001; Siaut et al., 2011; Merchant et al., 2012; Li-Beisson et al., 2015). Lipids possess crucial functions in many biological processes in plants and microalgae, such as maintaining cellular membrane structures, storing energy and carbons, and mediating signaling pathways (Guschina et al., 2006; Hu et al., 2008; Barbaglia and Hoffmann-Benning, 2016). Lipid profiles of algal cells may be affected by various environment stresses, such as high light and nutrient deprivation conditions (Solovchenko et al., 2008).
The traditional methods for lipids analysis include thin layer chromatography (TLC), gas chromatography (GC) and GC-mass spectrometry (GC/MS) (Diehl et al., 1995; Xu et al., 2002; Rainville et al., 2007). However, there are many disadvantages for these methods. TLC, GC, and GC/MS involve timeconsuming and laborious sample pre-treatment procedures. Moreover, these methods cannot be used to quantify the cellular content of a given lipid molecule, with limited applications in functional genomics studies (Rainville et al., 2007; CarrascoPancorbo et al., 2009).
In recent years, mass spectrometry (MS) based analytical methods have been used in analysis of lipidomes of biological samples with different genetic backgrounds or physiological interferences (Han and Gross, 2003, 2005; Welti et al., 2003; Rainville et al., 2007). Since microalgal glycerolipids have attracted great interest due to their potentials in energy and nutraceutical applications, lipidomes of a large number of microalgae including both membrane glycerolipids and triacylglycerols have been resolved (reviewed by De Costa et al., 2016). Lipidomics is a very effective biochemical analytical tool for precise identification of glycerollipids, which is not only able to resolve lipid classes and fatty acyl composition, but also can facilitate absolute quantification at molecular level, especially when coupled with ultra-high performance liquid chromatography that can essentially improve the sensitivity and reproducibility (Cutignano et al., 2016; Han et al., 2017; Řezanka et al., 2018).
For C. reinhardtii, the model species for dissecting lipid metabolism in microalgae, its lipidomes have been characterized in a few previous studies which showed monogalactosyldiacylglyceride (MGDG), digalactosydiacylglyceride (DGDG), sulfoquinovosyldiacylglycerol (SQDG), phosphatidylglycerol (PG), diacylglyceryl-N, N, N-trimethylhomoserine (DGTS), phosphatidylinositol (PI) and phosphatidyl ethanolamine (PE) are the major membrane lipids of C. reinhardtii (Yoon et al., 2012; Yang et al., 2015). MGDG, DGDG, SQDG, and PG constitute the chloroplast membranes in algal cells, playing important functional roles in photosynthesis (Harwood and Guschina, 2009; Kobayashi et al., 2016). In microalgae like C. reinhardtii, the extraplastidic membrane lipids are mainly composed of DGTS, PI and PE (Moellering et al., 2009), among which the betain lipid DGTS was the major class (Klug and Benning, 2001).
In this study, the analytical tool of ultra performance liquid chromatography (UPLC)-ESI-MS/MS was used to quantitatively analyze the lipidomes of C. reinhardtii wild type and starchless mutant sta6 cells. The starchless mutant sta6 was deficient of the gene encoding an ADP-glucose pyrophosphorylase subunit and hence incapable of synthesizing starch (Zabawinski et al., 2001). In previous study, shunting carbon precursors from the starch synthesis pathway in C. reinhardtii led to a 10-fold increase in triacylglycerol (TAG) content as compared to the wild type (Li et al., 2010a, b), but it is unknown whether inactivation of AGPase may have any effect on the cellular contents of membrane lipids. The study aims to investigate global changes in lipid metabolism and homeostasis in the starchless mutant. In addition, responses of such two C. reinardtii strains to the changing illumination conditions at the level of lipid molecules were investigated by using the lipidomics approach as well, which will underpin our understanding about the impact of blocking starch biosynthesis on cellular capabilities in adapting changing environmental conditions.
2 MATERIAL AND METHOD 2.1 Strains and growth conditionsChlamydomonas reinhardtii cc-124 (WT) was purchased from Chlamydomonas resource center at the University of Minnesota (https://www.chlamycollection.org), and sta6 was kindly provided by Arizona State University. Both strains were precultured in the TAP growth medium (Gorman and Levine, 1965) at 25℃ on an orbital shaker incubator (NBS Innova 44R, Eppendorf, Germany) at a constant rate of 100 r/min. The light intensity was kept at 50 μmol/(m2·s), which was measured by using a photosynthetically active radiation (PAR) meter (Mastercycler pro S, Eppendorf, Germany). Algal cells from the logarithmic phase were inoculated into the 800 mL column photobioreactor (i.d., 5 cm) containing 750 mL HSM medium (Sueoka, 1960), with an initial cell concentration of (2.0-2.5)×106 cells/mL. Algal cells were cultivated under 5 μmol/(m2·s) (low light, LL) for 3 days and then transferred to 150 μmol/(m2·s) (high light, HL) for 12 h. The algal samples from both illumination conditions were harvested by centrifugation at 3 000×g for 5 min at 4℃, and then freeze-dried.
2.2 Lipid extractionTotal lipids were extracted according to the method described previously (Wang and Benning, 2011) with minor modifications. The freeze-dried samples (about 10 mg, dry weight) were ground in liquid nitrogen and extracted with 6 mL solvents composed of methanol, chloroform and formic acid (20׃10׃1, v/v/v). The extracts were vigorously vortexed for 5 min followed by adding 3 mL of 0.2 mol/L H3PO4 and 1 mol/L KCl, and then vortexed briefly. After centrifuging at 1 000×g for 5 min, lipids dissolved in the lower chloroform phase were collected by using a glass pipette. Solvents were then evaporated under nitrogen stream. Lipid samples were stored at -80℃ prior to use.
2.3 Lipidomes analysis and quantification with ESI/MSLipidomes analyses were performed on a triple quadrupole MS/MS (Xevo TQ-S, Waters, USA) with electrospray ionization (ESI) source coupled with an Acquity Ultra-Performance Liquid Chromatography (UPLC) system (Waters, USA). Lipid samples were separated on a BEH C18 column (length 50 mm, internal diameter 2.1 mm, particle size 1.7 μm; Waters) prior to the ESI/MS analysis. The column temperature was kept at 30℃ and 35℃ for the positive and negative ion mode, respectively. The ESI/MS analysis was carried out according to the method developed by Yoon et al previously (Yoon et al., 2012) with modifications. Samples were recovered in 1 mL chloroform/methanol (1:1, v/v). For absolute quantification, lipids samples were mixed with internal standards (ISTD), including MGDG 18:0/18:0, DGDG 18:0/18:0, DGTS 16:0/16:0 d9, PE 17:0/14:1, PG 17:0/20:4 and PI 17:0/20:4. Among these ISTD, PG 17:0/20:4 was used for both PG and SQDG quantification. The external standards (ESTD) for calibration included MGDG mixture standard containing MGDG 16:3/18:3, MGDG 16:3/18:2 and MGDG 16:1/18:3, DGDG mixture standard containing DGDG 16:3/18:3, DGDG 16:3/18:2, DGDG 16:1/18:3 and DGDG 16:0/18:3, DGTS 16:0/16:0, PG 16:0/18:1, PG 18:0/18:1, PE 18:0/18:1, PI 18:1/18:1 and SQDG 16:0/18:3 were used as ETSDs for the corresponding classes of membrane lipids. Lipid standards were all purchased from Avanti Polar Lipids Ltd. (USA) except that MGDG 18:0/18:0 and DGDG 18:0/18:0 were obtained from Matreya LLC (USA). For quantification, ESTD were titrated relative to a constant amount of ISTD to establish the correlation between the ratio of the analyte signal to the ISTD signal and the ratio of their concentrations (Wang et al., 2014). MGDG, DGDG, and DGTS were detected in the positive mode, while PG, SQDG, PE, and PI were analyzed in the negative mode. Multiple reaction monitoring (MRM) was employed for quantitative analysis.
2.4 Data processingRaw files of the ESI/MS data were retrieved and analyzed with Masslynx v4.1 software (Waters). Student's t-test was used to compare the cellular contents of given lipids between WT and sta6 (n=4, including two biological replicates and two technical replicates). If the test gives P value ≤0.05, the differences between two samples were interpreted as being significant.
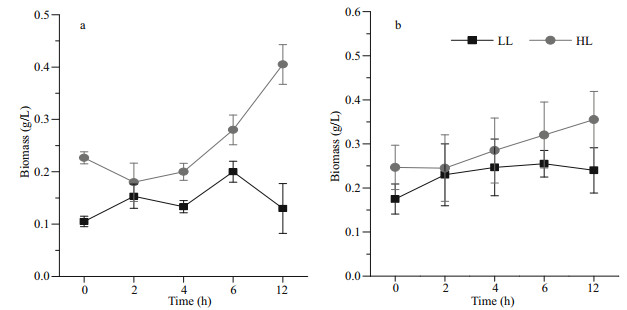
3 RESULT 3.1 Growth of cc-124 and sta6 under low light and high lightThe biomass of cc-124 increased slightly in the first 2 h and then was fluctuated over 12 h under the LL conditions (Fig. 1a). On the contrast, WT grew fast under the HL conditions, the biomass of which achieved 0.4 g/L after 12 h (Fig. 1a). Distinct from cc- 124, sta6 showed slight growth under two conditions and no significant difference was observed (Fig. 1b).
|
| Fig.1 Biomass production of WT (a) and sta6 (b) cultivated under low-light (LL) and high-light (HL) conditions |
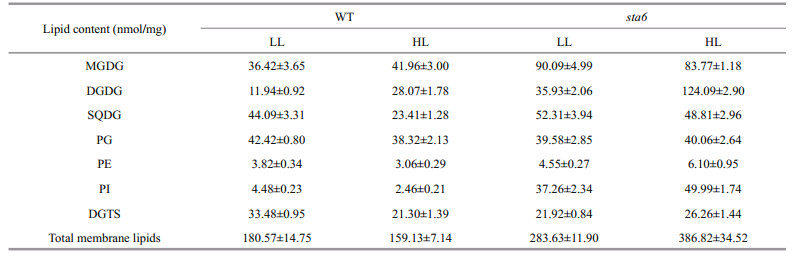
The lipidomics platform established in this study enabled absolute quantification of 7 classes of major membrane lipids of C. reinhardtii (Table 1). Under LL, the content of total membrane lipids was 180.57±14.75 and 283.63±11.90 nmol/mg (P < 0.001) for WT and sta6, respectively. Under HL, the content of total membrane lipids was reduced by 11.7% to 159.13±7.14 nmol/mg in WT, while it was increased by 36.4% and reached 386.82±34.52 nmol/mg in sta6.
|
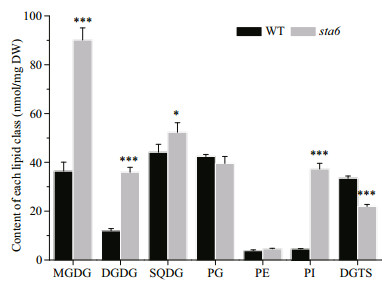
The galactolipids including MGDG and DGDG were the most abundant membrane lipids in WT and starchless mutant sta6 grown under LL, accounting for 27.38% and 44.74% of the total membrane lipids in them, respectively (Fig. 2, Table 1). In WT, the content of MGDG and DGDG was 36.42 nmol/mg DW and 11.94 nmol/mg DW, respectively. The content of MGDG and DGDG in sta6 was greater than that of WT by 147.35% (P < 0.001) and 200.78% (P < 0.001), respectively.
|
| Fig.2 Contents of membrane lipids in C. reinhardtii WT and sta6 cells under low light (LL) Values represent the mean ± SD (n=4). * P<0.05, *** P<0.001. |
Under LL, the most abundant MGDG molecular species was MGDG 18׃3/16:4 in WT, of which the content was 20.59 nmol/mg DW (Fig. 3a), accounting for 14.50% of the total MGDG. Different from WT, MGDG 18:1/16:1, 18:1/16:4, and 18:3/16:4 were the most abundant MGDG species in sta6 cells grown under LL, accounting for 25.77% (corresponding to 23.11 nmol/mg DW), 23.27% (20.86 nmol/mg DW), and 23.90% (21.43 nmol/mg DW) of its total MGDG pool, respectively. The major DGDG species in WT and sta6 under LL was both DGDG 18:3/16:1, which was 7.6 and 30.87 nmol/mg DW (Fig. 3b), corresponding to 65.07% and 86.78% of the total DGDG, respectively.

|
| Fig.3 Contents of chloroplast membrane lipids molecular species in C. reinhardtii WT and sta6 under low light (LL) a. MGDG; b. DGDG; c. SQDG; d. PG. Values represent the mean±SD (n=4). *: P<0.05; **: P<0.01; ***: P<0.001. |
SQDG and PG were two types of polar lipids present in the chloroplast membranes. The total SQDG content in sta6 under LL was 53.92 nmol/mg DW, which was 18.64% higher than in WT (44.09 nmol/mg DW, P < 0.05) (Fig. 2). Among SQDG molecular species, the content of SQDG 16:0/16:0 was 38.20 and 38.29 nmol/mg DW in WT and sta6, respectively (Fig. 3c), which was the major species in both WT and sta6. Among SQDG molecules, one of the highly unsaturated species SQDG 16:0/18:4 was the one enhanced to the most extent, which was increased by 975.84% (P < 0.001) in sta6 as compared with WT. As for PG, the only bulk phospholipid found in chloroplast (Wada and Murata, 2007), WT and sta6 contained similar amounts of them, of which the cellular content was 42.42 and 39.58 nmol/mg DW, respectively (Fig. 2). The most abundant PG species were PG 16:1/18:3 in WT, whereas it was replaced by PG 16:1/18:1 in sta6 (Fig. 3d), which accounted for 41.09% and 41.81% of their total PG, respectively. In addition, PG 16:1/18:1 showed the greatest change with an increase by 287.80% (P < 0.001) in sta6 compared to WT.
PI and PE were two types of phospholipids constituting the extraplastidic membranes in C. reinhardtii. Under LL, the total PI content was 4.36 nmol/mg DW in WT (Fig. 2), among which PI 16:0/18:1 was the major species accounting for 94.24% of the total PI (Fig. 4a). In sta6, the content of PI was increased dramatically by 731.58% (P < 0.001) compared to that of WT. There was no significant change in total PE content between WT and sta6, which were 3.82 and 4.55 nmol/mg DW, respectively (Fig. 2). Two PE species including 18:0/18:3 and 18:2/18:3 were identified in both strains, among which PE 18:0/18:3 was the major species (Fig. 4a).

|
| Fig.4 Contents of extraplastidic membrane lipids molecular species in C. reinhardtii WT and sta6 under low light (LL) a. PI and PE; b. DGTS major species; c. DGTS minor species. Values represent the mean±SD (n=4). *: P<0.05; **: P<0.01; ***: P<0.001. |
A large number of DGTS molecular species were identified in both strains. DGTS 16:0/18:3 was the most abundant species in WT (7.61 nmol/mg DW), whereas DGTS 16:0/16:1, 16:0/18:3 and 16:0/18:4 were three species with the highest contents in sta6, which was 3.08, 3.9, and 4.27 nmol/mg DW, respectively (Fig. 4b). Among the membrane lipids, DGTS was the only group of lipid class that was reduced in sta6 compared with WT (Fig. 2, 33.48 vs 21.92 nmol/mg DW, P < 0.001).
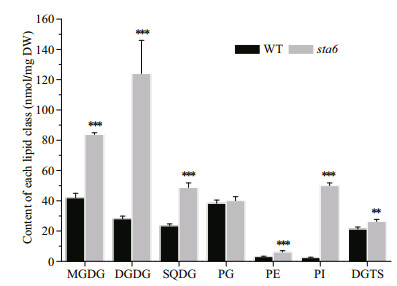
3.3 Lipidomes of cc-124 and sta6 cells grown under high lightAfter high light acclimation for 12 h, the contents of all the membrane lipids were significantly increased in sta6 as compared to WT, except for PG. The chloroplast membrane lipid MGDG, DGDG, and SQDG was increased by 99.62% (P < 0.001), 342.14% (P < 0.001) and 108.50% (P < 0.001), respectively, while the extraplastidic membrane lipids PI, PE, and DGTS was increased by 19-fold (P < 0.001), 99.35% (P < 0.001) and 23.32% (P < 0.01), respectively (Fig. 5).
|
| Fig.5 Contents of membrane lipids in C. reinhardtii WT and sta6 cells under high light (HL) Values represent the mean±SD (n=4). **: P<0.01; ***: P<0.001. |
The cellular contents of the chloroplast membrane lipid molecules were shown in Fig. 6. For galactolipids, most MGDG and DGDG species in sta6 were higher than that in WT (Fig. 6a & b). The cellular content of MGDG 18:1/16:3 in WT was 0.14 nmol/mg DW, whereas it was 4.96 nmol/mg DW in sta6, increased by 33-fold (Fig. 6a, P < 0.001). The predominant DGDG 18:3/16:1 was 66.55 nmol/mg DW in sta6, upregulated by 392.51% compared to WT (Fig. 6b, 13.51 nmol/mg DW, P < 0.001). Similar to galactolipids, most SQDG species were up-regulated in sta6 compared to in WT under HL. Among SQDG species, the most abundant SQDG 16:0/18:0 changed dramatically from 0.14 nmol/mg DW in WT to 0.63 nmol/mg DW in sta6, increased by 364.67% (Fig. 6c, P < 0.001). The total PG in two strains showed no significant change (Fig. 5), because the cellular contents of two predominant species PG 16:1/18:2 and PG 16:1/18:3 were unchanged (Fig. 6d). However, several minor PG species were increased in sta6 compared to WT, including PG 16:1/18:1 increased by 138.63% (P < 0.01) and PG 16:0/16:1 decreased by 41.41% (P < 0.01) in sta6 as compared with WT (Fig. 6d).

|
| Fig.6 Contents of chloroplast membrane lipids molecular species in C. reinhardtii WT and sta6 under high light (HL) a. MGDG; b. DGDG; c. SQDG; d. PG. Values represent the mean±SD (n=4). *: P<0.05; **: P<0.01; ***: P<0.001. |
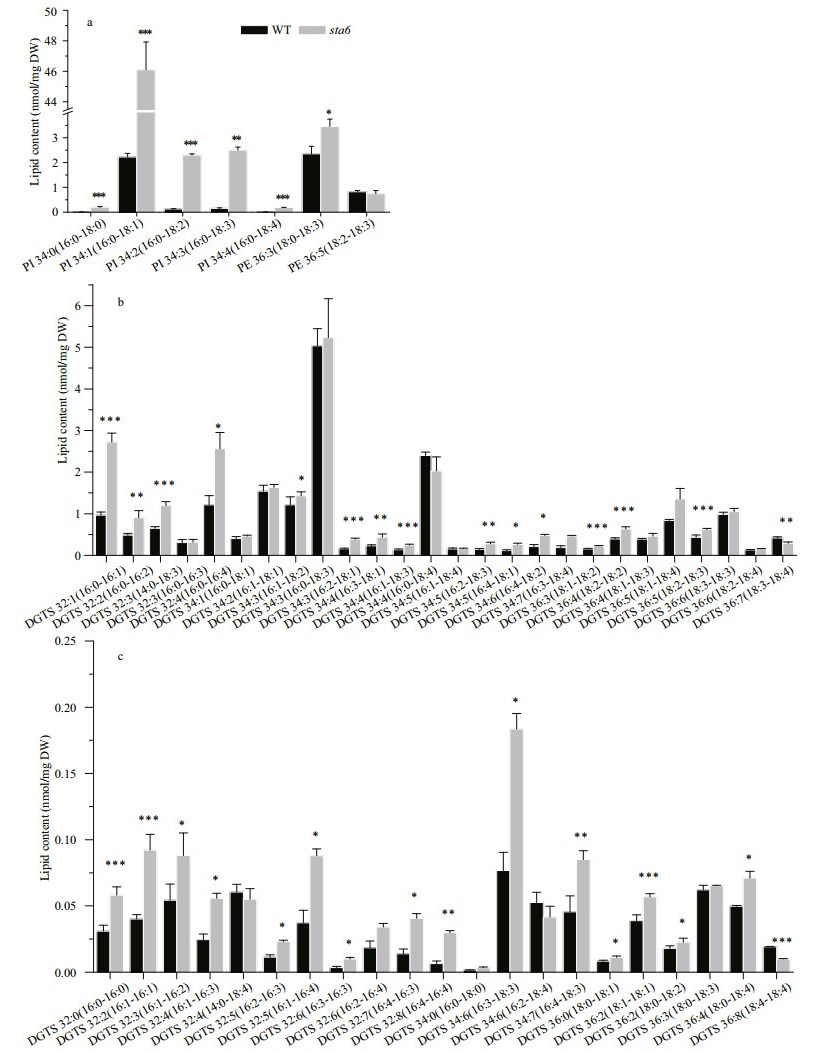
Similar to the drastic changes in the chloroplast membrane lipids, the cellular content of extraplastidic membrane lipids including PI, PE, and DGTS showed drastic increase in sta6 by 19-fold (P < 0.001), 99.35% (P < 0.001), and 23.32% (P < 0.01), respectively, as compared to that in WT (Fig. 7). Each PI molecular species in sta6 was increased compared to that of WT under HL, among which the highly saturated PI 16:0/18:0 showed the largest increase by 22-fold (P < 0.001) (Fig. 7a). For the two PE species that identified in both WT and sta6, PE 18:0/18:3 rapidly increased by 46.84% in sta6 as compared with that in WT (Fig. 7a, 2.35 nmol/mg DW, P < 0.05), whereas PE 18:2/18:3 remained unchanged. In numerous DGTS species, DGTS 16:0/16:1 increased to the most in sta6 (Fig. 7b, P < 0.001), which was 182.70% higher than that of WT. However, a large number of DGTS species remained relatively stable in both strains, such as the dominant DGTS species, DGTS 16:0/18:3 (Fig. 7b). By contrast, two DGTS species with highly unsaturated fatty acids, DGTS 18:3/18:4 and DGTS 18:4/18:4, were reduced by 31.33% (Fig. 7b, P < 0.01), and 47.93% (Fig. 7c, P < 0.001) in sta6, respectively, when compared to that of WT.
|
| Fig.7 Contents of extraplastidic membrane lipids molecular species in C. reinhardtii WT and sta6 under high light (HL) a. PI and PE; b. DGTS major species; c. DGTS minor species. Values represent the mean±SD (n=4). *: P<0.05; **: P<0.01; ***: P<0.001. |
The starchless mutant sta6 has been intensively investigated due to its capabilities in overaccumulating triacylglycerols under stress conditions (Krishnan et al., 2015; Fan et al., 2017; Tran et al., 2019). However, mechanisms underlying enhanced TAG biosynthesis in sta6 remained elusive. There are emerging studies showing sta6 exhibited comprehensive phenotypes, including retarded cellular growth under certain conditions, increased central carbon metabolism, and slowed NADPH reoxidation (Blaby et al., 2013; Krishnan et al., 2015). This study aimed at dissecting the alterations in the cellular structures with respect to the membrane lipids compositions, which will provide insights into the physiological consequences of blocking starch biosynthesis in microalgae.
By using the ESI/MS method, a total of 16 MGDG molecules, 16 DGDG molecules, 6 SQDG molecules, 11 PG molecules, 5 PI molecules, 2 PE molecules and 49 DGTS molecules were identified in both C. reinhardtii WT and sta6, indicative of that the membrane profiles are very similar between these two strains. On the other hand, quantitative analysis revealed that the cellular contents of a large number of lipid molecules were changed in sta6, due to the deficiency in starch synthesis.
In general, the chloroplast membranes of C. reinhardtii are composed of the same lipid classes as the chloroplasts of plants and algae (Goss and Wilhelm, 2009). The chloroplast membranes of C. reinhardtii are enriched in uncharged galactolipids, MGDG and DGDG (Riekhof et al., 2003), corresponding to 60%–80% of total chloroplast lipids in oxygenic photosynthetic organisms and MGDG is the predominant one (Block et al., 1983; Riekhof et al., 2003; Sakurai et al., 2006). However, in this study, although galactolipids MGDG and DGDG are still the prevalent chloroplast lipids in both C. reinhardtii strains under LL and HL, DGDG became dominant in sta6 under HL (Fig. 5). This result suggests that the MGDG/DGDG ratio was somehow changed in starchless mutant, probably to maintain the structure and stability of the thylakoid membranes under HL (Murphy, 1982; Demé et al., 2014), because DGDG is a typical bilayer forming lipids while MGDG tended non-lamellar and hexagonal structures due to its conelike molecular shape (Sen et al., 1981; Webb and Green, 1991).
Starch and lipids are two major energy storage forms in most unicellular green algae, especially under many stress conditions, such as high light. When the starch synthesis pathway was switched off, the metabolic shift of carbon flux away from starch synthesis into lipids synthesis pathways (Li et al., 2010b). It was found in this study that most chloroplast membrane lipids, except PG, showed higher concentrations in sta6 than in WT under different light conditions, especially under HL. These results indicated that the formation of more thylakoid membranes and photosynthetic complexes might have occurred in the starchless mutant. Similar to two diatoms wild type strains Cyclotella meneghiniana and Phaeodactylum tricornutum (Lepetit et al., 2012), illumination of sta6 with HL leads to a further increase of the concentration of anionic lipid SQDG, as compared with WT.
The betaine lipid DGTS have been identified as the substitutes for phosphatidylcholine (PC) constituting the extraplastidic membranes in many green algae including C. reinhardtii (Dembitsky, 1996). However, we found that PI replaced DGTS and became the major extraplastidic lipids in sta6. Since PI can be phosphorylated into phosphatidylinositol phosphates, such as phosphatidylinositol 4, 5-bisphospate (PI(4, 5) P2), participating in signaling pathways (Boss and Im, 2012), we assumed the accumulated PI may play important role in signaling in sta6 to cope with profoundly changed metabolism caused by blocked starch biosynthesis.
5 CONCLUSIONTo better understand the impact of blocking starch synthesis on lipid metabolism and homeostasis of C. reinhardtii, we have employed a UPLC-ESI-MS/MS method to qualitatively and quantitatively analyze the membrane lipids of WT and starchless mutant sta6. The results indicated that the lipid profile of sta6 is similar to that of WT, but cellular content of lipids are dramatically different between these two strains. In sta6, a large number of membrane lipids were upregulated under both LL and HL conditions, which indicated a great amount of photosynthetically-fixed carbons were shunt into membrane lipid biosynthesis due to blockage of starch biosynthesis.
6 DATA AVAILABILITY STATEMENTAll data generated and/or analyzed during this study are included in this published article.
Barbaglia A M, Hoffmann-Benning S. 2016. Long-distance lipid signaling and its role in plant development and stress response. In: Nakamura Y, Li-Beisson Y eds. Lipids in Plant and Algae Development. Springer, Cham. p.339-361.
|
Blaby I K, Glaesener A G, Mettler T, Fitz-Gibbon S T, Gallaher S D, Liu B S, Boyle N R, Kropat J, Stitt M, Johnson S, Benning C, Pellegrini M, Casero D, Merchant S S. 2013. Systems-level analysis of nitrogen starvation-induced modifications of carbon metabolism in a Chlamydomonas reinhardtii starchless mutant. The Plant Cell, 25(11): 4305-4323.
DOI:10.1105/tpc.113.117580 |
Block M A, Dorne A J, Joyard J, Douce R. 1983. Preparation and characterization of membrane fractions enriched in outer and inner envelope membranes from spinach chloroplasts.II. Biochemical characterization. Journal of Biological Chemistry, 258(21): 13273-13286.
|
Boss W F, Im Y J. 2012. Phosphoinositide signaling. Annual Review of Plant Biology, 63(1): 409-429.
DOI:10.1146/annurev-arplant-042110-103840 |
Carrasco-Pancorbo A, Navas-Iglesias N, Cuadros-Rodríguez L. 2009. From lipid analysis towards lipidomics, a new challenge for the analytical chemistry of the 21st century.Part I:Modern lipid analysis. TrAC Trends in Analytical Chemistry, 28(3): 263-278.
DOI:10.1016/j.trac.2008.12.004 |
Cutignano A, Luongo E, Nuzzo G, Pagano D, Manzo E, Sardo A, Fontana A. 2016. Profiling of complex lipids in marine microalgae by UHPLC/tandem mass spectrometry. Algal Research, 17: 348-358.
DOI:10.1016/j.algal.2016.05.016 |
Da Costa E, Silva J, Mendonça S, Abreu M, Domingues M. 2016. Lipidomic approaches towards deciphering glycolipids from microalgae as a reservoir of bioactive lipids. Marine Drugs, 14(5): 101.
DOI:10.3390/md14050101 |
Dembitsky V M. 1996. Betaine ether-linked glycerolipids:chemistry and biology. Progress in Lipid Research, 35(1): 1-51.
DOI:10.1016/0163-7827(95)00009-7 |
Demé B, Cataye C, Block M, Maréchal E, Jouhet J. 2014. Contribution of galactoglycerolipids to the 3-dimensional architecture of thylakoids. The FASEB Journal, 28(8): 3373-3383.
DOI:10.1096/fj.13-247395 |
Diehl B W K, Herling H, Riedl I, Heinz E. 1995. 13C-NMR analysis of the positional distribution of fatty acids in plant glycolipids. Chemistry and Physics of Lipids, 77(2): 147-153.
DOI:10.1016/0009-3084(95)02462-r |
Fan J H, Zheng L H, Bai Y P, Saroussi S, Grossman A R. 2017. Flocculation of Chlamydomonas reinhardtii with different phenotypic traits by metal cations and high pH. Frontiers in Plant Science, 8: 1997.
DOI:10.3389/fpls.2017.01997 |
Gorman D S, Levine R P. 1965. Cytochrome f and plastocyanin:their sequence in the photosynthetic electron transport chain of Chlamydomonas reinhardi. Proceedings of the National Academy of Sciences of the United States of America, 54(6): 1665-1669.
DOI:10.1073/pnas.54.6.1665 |
Goss R, Wilhelm C. 2009. Lipids in algae, lichens and mosses.In: Wada H, Murata N eds. Lipids in Photosynthesis.Springer, Dordrecht. p.117-137.
|
Guschina I A, Harwood J L. 2006. Lipids and lipid metabolism in eukaryotic algae. Progress in Lipid Research, 45(2): 160-186.
DOI:10.1016/j.plipres.2006.01.001 |
Han D X, Jia J, Li J, Sommerfeld M, Xu J, Hu Q. 2017. Metabolic remodeling of membrane glycerolipids in the microalga Nannochloropsis oceanica under nitrogen deprivation. Frontiers in Marine Science, 7: 402.
DOI:10.3389/fmars.2017.00242 |
Han X L, Gross R W. 2003. Global analyses of cellular lipidomes directly from crude extracts of biological samples by ESI mass spectrometry:a bridge to lipidomics. Journal of Lipid Research, 44(6): 1071-1079.
DOI:10.1194/jlr.R300004-JLR200 |
Han X L, Gross R W. 2005. Shotgun lipidomics:electrospray ionization mass spectrometric analysis and quantitation of cellular lipidomes directly from crude extracts of biological samples. Mass Spectrometry Reviews, 24(3): 367-412.
DOI:10.1002/mas.20023 |
Harris E H. 2001. Chlamydomonas as a model organism. Annu Rev Plant Phys, 52(1): 363-406.
DOI:10.1146/annurev.arplant.52.1.363 |
Harwood J L, Guschina I A. 2009. The versatility of algae and their lipid metabolism. Biochimie, 91(6): 679-684.
DOI:10.1016/j.biochi.2008.11.004 |
Hu Q, Sommerfeld M, Jarvis E, Ghirardi M, Posewitz M, Seibert M, Darzins A. 2008. Microalgal triacylglycerols as feedstocks for biofuel production:perspectives and advances. The Plant Journal, 54(4): 621-639.
DOI:10.1111/j.1365-313X.2008.03492.x |
Klug R M, Benning C. 2001. Two enzymes of diacylglycerylO-4'-(N, N, N, -trimethyl) homoserine biosynthesis are encoded by btaA and btaB in the purple bacterium Rhodobacter sphaeroides. Proceedings of the National Academy of Sciences of the United States of America, 98(10): 5910-5915.
DOI:10.1073/pnas.101037998 |
Kobayashi K, Endo K, Wada H. 2016. Roles of lipids in photosynthesis. In: Nakamura Y, Li-Beisson Y eds. Lipids in Plant and Algae Development. Springer, Cham. p.21-49.
|
Krishnan A, Kumaraswamy G K, Vinyard D J, Gu H Y, Ananyev G, Posewitz M C, Dismukes G C. 2015. Metabolic and photosynthetic consequences of blocking starch biosynthesis in the green alga Chlamydomonas reinhardtii sta6 mutant. The Plant Journal, 81(6): 947-960.
DOI:10.1111/tpj.12783 |
Lepetit B, Goss R, Jakob T, Wilhelm C. 2012. Molecular dynamics of the diatom thylakoid membrane under different light conditions. Photosynthesis Research, 111(1-2): 245-257.
DOI:10.1007/s11120-011-9633-5 |
Li Y T, Han D X, Hu G R, Dauvillee D, Sommerfeld M, Ball S, Hu Q. 2010a. Chlamydomonas starchless mutant defective in ADP-glucose pyrophosphorylase hyperaccumulates triacylglycerol. Metabolic Engineering, 12(4): 387-391.
DOI:10.1016/j.ymben.2010.02.002 |
Li Y T, Han D X, Hu G R, Sommerfeld M, Hu Q. 2010b. Inhibition of starch synthesis results in overproduction of lipids in Chlamydomonas reinhardtii. Biotechnology and Bioengineering, 107(2): 258-268.
DOI:10.1002/bit.22807 |
Li-Beisson Y, Beisson F, Riekhof W. 2015. Metabolism of acyl-lipids in Chlamydomonas reinhardtii. The Plant Journal, 82(3): 504-522.
DOI:10.1111/tpj.12787 |
Merchant S S, Kropat J, Liu B S, Shaw J, Warakanont J. 2012. TAG, you're it! Chlamydomonas as a reference organism for understanding algal triacylglycerol accumulation. Current Opinion in Biotechnology, 23(3): 352-363.
DOI:10.1016/j.copbio.2011.12.001 |
Moellering E R, Miller R, Benning C. 2009. Molecular genetics of lipid metabolism in the model green alga Chlamydomonas reinhardtii. In: Wada H, Murata N eds.Lipids in Photosynthesis. Springer, Dordrecht. p.139-155.
|
Murphy D J. 1982. The importance of non-planar bilayer regions in photosynthetic membranes and their stabilisation by galactolipids. FEBS Letters, 150(1): 19-26.
DOI:10.1016/0014-5793(82)81297-0 |
Řezanka T, Lukavský J, Vítová M, Nedbalová L, Sigler K. 2018. Lipidomic analysis of Botryococcus(Trebouxiophyceae, Chlorophyta)-Identification of lipid classes containing very long chain fatty acids by offline two-dimensional LC-tandem MS. Phytochemistry, 148: 29-38.
DOI:10.1016/j.phytochem.2018.01.011 |
Riekhof W R, Ruckle M E, Lydic T A, Sears B B, Benning C. 2003. The sulfolipids 2'-O-acylsulfoquinovosyldiacylglycerol and sulfoquinovosyldiacylglycerol are absent from a Chlamydomonas reinhardtii mutant deleted in SQD1. Plant Physiology, 133(2): 864-874.
DOI:10.1104/pp.103.029249 |
Sakurai I, Shen J R, Leng J, Ohashi S, Kobayashi M, Wada H. 2006. Lipids in oxygen-evolving photosystem II complexes of cyanobacteria and higher plants. The Journal of Biochemistry, 140(2): 201-209.
DOI:10.1093/jb/mvj141 |
Sen A, Williams W P, Quinn P J. 1981. The structure and thermotropic properties of pure 1, 2-diacylgalactosylglycerols in aqueous systems. Biochimica et Biophysica Acta (BBA)-Lipids and Lipid Metabolism, 663(2): 380-389.
DOI:10.1016/0005-2760(81)90167-3 |
Siaut M, Cuiné S, Cagnon C, Fessler B, Nguyen M, Carrier P, Beyly A, Beisson F, Triantaphylidès C, Li-Beisson Y, Peltier G. 2011. Oil accumulation in the model green alga Chlamydomonas reinhardtii:characterization, variability between common laboratory strains and relationship with starch reserves. BMC Biotechnology, 11: 7.
DOI:10.1186/1472-6750-11-7 |
Solovchenko A E, Khozin-Goldberg I, Didi-Cohen S, Cohen Z, Merzlyak M N. 2008. Effects of light intensity and nitrogen starvation on growth, total fatty acids and arachidonic acid in the green microalga Parietochloris incisa. Journal of Applied Phycology, 20(3): 245-251.
DOI:10.1007/s10811-007-9233-0 |
Sueoka N. 1960. Mitotic replication of deoxyribonucleic acid in Chlamydomonas reinhardi. Proceedings of the National Academy of Sciences of the United States of America, 46(1): 83-91.
DOI:10.1073/pnas.46.1.83 |
Tran Q G, Cho K, Park S B, Kim U, Lee Y J, Kim H S. 2019. Impairment of starch biosynthesis results in elevated oxidative stress and autophagy activity in Chlamydomonas reinhardtii. Scientific Reports, 9: 9856.
DOI:10.1038/s41598-019-46313-6 |
UPLC-oaTOF-MS for lipid analysis in complex biological mixtures: A new tool for lipidomics. Journal of Proteome Research, 6(2): 552-558, https://doi.org/10.1021/pr060611b.
|
Wada H, Murata N. 2007. The essential role of phosphatidylglycerol in photosynthesis. Photosynthesis Research, 92(2): 205-215.
DOI:10.1007/s11120-007-9203-z |
Wang B B, Zhang Z, Hu Q, Sommerfeld M, Lu Y H, Han D X. 2014. Cellular capacities for high-light acclimation and changing lipid profiles across life cycle stages of the green alga Haematococcus pluvialis. PLoS One, 9(9): e106679.
DOI:10.1371/journal.pone.0106679 |
Wang Z, Benning C. 2011. Arabidopsis thaliana polar glycerolipid profiling by thin layer chromatography(TLC) coupled with gas-liquid chromatography (GLC). Journal of Visualized Experiments, 49(2): 518.
DOI:10.3791/2518 |
Webb M S, Green B R. 1991. Biochemical and biophysical properties of thylakoid acyl lipids. Biochimica et Biophysica Acta (BBA)-Bioenergetics, 1060(2): 133-158.
DOI:10.1016/S0005-2728(09)91002-7 |
Welti R, Wang X M, Williams T D. 2003. Electrospray ionization tandem mass spectrometry scan modes for plant chloroplast lipids. Analytical Biochemistry, 314(1): 149-152.
DOI:10.1016/S0003-2697(02)00623-1 |
Xu Y N, Wang Z N, Yan X J, Lin W, Li L B, Kuang T Y. 2002. Positional distribution of fatty acids on the glycerol backbone during the biosynthesis of glycerolipids in Ectocarpus fasciculatus. Chinese Science Bulletin, 47(21): 1802-1806.
DOI:10.1007/BF03183846 |
Yang D W, Song D H, Kind T, Ma Y, Hoefkens J, Fiehn O. 2015. Lipidomic analysis of Chlamydomonas reinhardtii under nitrogen and sulfur deprivation. PLoS One, 10(9): e0137948.
DOI:10.1371/journal.pone.0137948 |
Yoon K, Han D X, Li Y T, Sommerfeld M, Hu Q. 2012. Phospholipid:diacylglycerol acyltransferase is a multifunctional enzyme involved in membrane lipid turnover and degradation while synthesizing triacylglycerol in the unicellular green microalga Chlamydomonas reinhardtii. The Plant Cell, 24(9): 3708-3724.
DOI:10.1105/tpc.112.100701 |
Zabawinski C, Van Den Koornhuyse N, D'Hulst C, Schlichting R, Giersch C, Delrue B, Lacroix J M, Preiss J, Ball S. 2001. Starchless mutants of Chlamydomonas reinhardtii lack the small subunit of a heterotetrameric ADP-glucose pyrophosphorylase. Journal of Bacteriology, 183(3): 1069-1077.
DOI:10.1128/jb.183.3.1069-1077.2001 |
 2020, Vol. 38
2020, Vol. 38


