Institute of Oceanology, Chinese Academy of Sciences
Article Information
- LI Mengna, CHEN Hao, WANG Minxiao, ZHONG Zhaoshan, ZHOU Li, LI Chaolun
- Identification and characterization of endosymbiosis-related immune genes in deep-sea mussels Gigantidas platifrons
- Journal of Oceanology and Limnology, 38(4): 1292-1303
- http://dx.doi.org/10.1007/s00343-020-0040-7
Article History
- Received Jan. 17, 2020
- accepted in principle Mar. 27, 2020
- accepted for publication May. 18, 2020
2 Laboratory for Marine Ecology and Environmental Science, Qingdao National Laboratory for Marine Science and Technology, Qingdao 266237, China;
3 Center for Ocean Mega-Science, Chinese Academy of Sciences, Qingdao 266071, China;
4 University of Chinese Academy of Sciences, Beijing 100049, China
Deep-sea cold seeps and hydrothermal vents are characterized by darkness, high-pressure and high concentration of sulfur and methane (Corliss et al., 1979; Kennicutt II et al., 1985). However, flourished chemosynthesis-driven ecosystems where chemolithoautotrophs act as primary producers have been continuingly found in these harsh environments (Sievert and Vetriani, 2015). Besides these chemolithoautotrophs living on hydrogen sulfide and methane, diversity of invertebrates including mussels, squat lobsters and tubeworms were also found thriving there with high biomass. Moreover, some of them have evolved with specific strategies to cope with the extreme environments (Brooks et al., 1987; Danovaro et al., 2014, 2017). And among these strategies, the intimate symbiosis with chemosynthetic microorganisms is of most interest and suggested as the key trait to thrive in deep-sea extreme environments (Van Dover et al., 2002).
Bathymodiolin mussels are common and numerically dominant macrofauna widely distributed in cold seeps and hydrothermal vents (Sibuet and Olu, 1998). They could obtain most of nutrients needed from methane-oxidizing bacteria (MOB) and/or sulfide-oxidizing bacteria (SOB) that are hosted in specialized epidermal cells of their gills, bacteriocytes (Dubilier et al., 2008). Besides, the symbiotic relationship could also shelter both hosts and symbionts from other environmental stresses (Sayavedra et al., 2015; Ponnudurai et al., 2017). Given their biological and ecological characteristics, Bathymodiolin mussels have therefore been regarded as a model organism for investigating the symbiotic interaction of deep-sea macrofauna and chemosynthetic microorganisms, as well as the adaptation of invertebrates in deep-sea. Meanwhile, the biological processes underlying the symbiosis have been clearer due to massive efforts made in recent years. For example, it was found that the chemosynthetic symbionts could be acquired from the surrounding environment through horizontal transmission during the juvenile stage of Bathymodiolin mussels (Fontanez and Cavanaugh, 2014). Meanwhile, an epithelial-wide distribution of symbionts could also be observed in juvenile mussels instead of exclusive distribution in gill bacteriocytes of adult hosts (Won et al., 2003; Wentrup et al., 2013). After the establishment of symbiosis, it was suggested that host mussels can obtain nutrients from either directly digesting symbionts via lysosome ("farming" hypothesis) (Fiala-Médioni et al., 2002) or indirectly assimilating the metabolic products secreted by symbionts ("milking" hypothesis) (Kádár et al., 2008). Nevertheless, molecules involved in above processes have been largely uncovered to date.
Gigantidas platifrons (previously referred to as Bathymodiolus platifrons) is a widely distributed species in the hydrothermal vents and methane seeps of Western Pacific Ocean (Fujiwara et al, 2000; Barry et al., 2002; Feng et al., 2015). As known, G. platifrons can only harbor methane-oxidizing bacteria in bacteriocytes of gills (Barry et al., 2002). The specific and robust symbiosis between G. platifrons and MOBs is suitable for studying how hosts interact with symbionts in deep-sea. As known, the selective establishment of symbiosis between hosts and symbionts depends largely on the innate immunity. The immune recognition of symbionts mediated by pattern recognition receptors (PRRs) could be the first step of symbiosis (Chow et al., 2010; Chu and Mazmanian, 2013). Moreover, endocytosis or phagocytosis of bacteria could also play an important role in the establishment of intracellular symbiosis. Recently, multiple immune related genes have been found positively selected or highly expressed in deep-sea mussels, suggesting their participation in symbiosis (Bettencourt et al., 2009, 2010; Zheng et al., 2017). However, none of them were verified or further characterized due to noncultivable symbionts and difficulties in sampling. In the present study, lipopolysaccharides (LPS) pull-down assay along with in situ hybridization (ISH) were conducted to isolate G. platifrons proteins potentially involved in the symbiosis with symbiotic methane oxidation bacteria. Moreover, possible roles of above genes were also investigated for better understanding how symbiosis was established and maintained.
2 MATERIAL AND METHOD 2.1 Mussels collection and preparationThe G. platifrons mussels were collected from the Formosa ridge cold seep of the South China Sea (22°06′N, 119°17′E), using remotely operated vehicle (ROV) Faxian (Discovery in Chinese) operated from R/V Kexue during cruise 2017 dive 149. Mussels were transferred to the deck with thermo-insulated Biobox to avoid temperature variation. Once onboard the R/V Kexue (Science in Chinese) mussel gills from fifty individuals were immediately dissected and stored in liquid nitrogen for pull-down assay. For ISH, freshly collected gills were fixed in cold 4% paraformaldehyde at 4℃ overnight. The gills were washed 3 times in phosphate buffer saline (PBS) and dehydrated in 75% ethanol, and stored in -20℃ before use.
2.2 LPS pull-down assayGiven that symbiotic MOBs are Gram-negative bacteria with LPS structure and cannot be cultured in vitro, LPS pull-down assay was conducted to isolate host proteins that potentially binding with symbionts according to the method reported previously (Xu et al., 2016). Briefly, 5 mg/mL LPS was firstly coupled to an Epoxy-activated Sepharose 6B column. After washing away unbound LPS, a total of 3-mg gill proteins (1 mg/mL in PBS with phenylmethanesulfonyl fluoride (PMSF)) were added to the column and incubated overnight. Subsequently, the column was thoroughly washed with PBS buffer to remove nonspecifically bound gill proteins, and then the protein samples were eluted with LPS (7.5 mg/mL) and urea (8 mol/L). Host gill proteins binding with LPS were finally separated by sodium dodecyl sulfatepolyacrylamide gel electrophoresis (SDS-PAGE) and further characterized by lipid chromatographytandem mass spectrometry (LC-MS/MS). Protein annotation was conducted given genome information released before (Wong et al., 2015; Sun et al., 2017) and updated by our lab (unpublished data).
2.3 Gene cloning and bioinformatics analysisFor ISH, full-length cDNA of vacuolar sorting protein (VSP), toll-like receptor 2 (TLR2), and leucine-rich repeat (LRR-1) were cloned using cDNA template from gill tissues, and the primers were designed based on the genome information of G. platifrons (Table 1). The PCR reactions were performed using exTaq (TaKaRa) according to the manual. The PCR program was set as follow: 94℃ for 5 min, 35 cycles of 94℃ for 20 s, 50-65℃ (determined by Tm of primers) for 30 s and 72℃ for 30 s–1 min (1 min of extension time per 1 kb amplicon), followed by the final extension at 72℃ for 10 min. The PCR products were purified and then connected into the pMD19-T vector for sequence verification. Full-length cDNA was translated into protein sequence and subjected to protein motif and structure prediction. The motif features were predicted by SMART (http://smart.embl-heidelberg.de) and the presumed tertiary structure of identified proteins was conducted by Swiss-Model (http://swissmodel.expasy.org/interactive). The signal peptides were predicted by SignalP-5.0 (http://www.cbs.dtu.dk/services/SignalP/).
Based on the results of pull-down experiments, VSP, TLR2, and LRR-1 genes were selected as representatives and subjected to ISH to investigate their expression patterns in gills. The synthesis of ISH probes and ISH assay were conducted using the method described by Halary et al. (2008). In brief, the DNA fragments of above genes were first amplified using specific primers pair (Table 1) as probe templates. Digoxigenin-labeled ssRNA probes including antisense probes and sense probes were synthesized using DIG RNA Labeling kit (Roche). Meanwhile, gills stored in 75% ethanol were dehydrated and embedded with paraffin. After sectioned at 7 μm thickness, gill samples were firstly treated with 10 μg/mL proteinase K for 10 min at 37℃ to increase permeability, and pre-hybridized in hybridization buffer (50% formamide, 5×saline sodium citrate (SSC), 50 μg/mL Heparin sodium, 0.5 mg/mL fish sperm DNA, and 0.01% Tween 20) for 3 h at 37℃. Subsequently, gill sections were hybridized for 12–16 h at 37℃ with 0.5 ng/μL probes in hybridization buffer. Afterward, the slides were washed in phosphate buffered saline with Tween 20 (PBST) and 4×SSC twice for 10 min each, and incubated with 2% bovine serum albumin (BSA) for 30 min along with 1‰ anti-digoxigenin-AP Fab fragments for 4 h. After washed by PBST, sections were incubated in development buffer pH 9.5 (100 mmol/L Tris-HCl, 100 mmol/L NaCl, 50 mmol/L MgCl2) for 10 min and stained with 5-bromo-4- chloro-3-indolyl-phosphate/nitro-blue-tetrazolium (BCIP/NBT) for 6–12 h. The reaction was then terminated by 0.5 mol/L ethylenediaminetetraacetic acid (EDTA) and imaged with microscopy after staining nucleic acids with 4', 6-diamidino-2- phenylindole (DAPI).
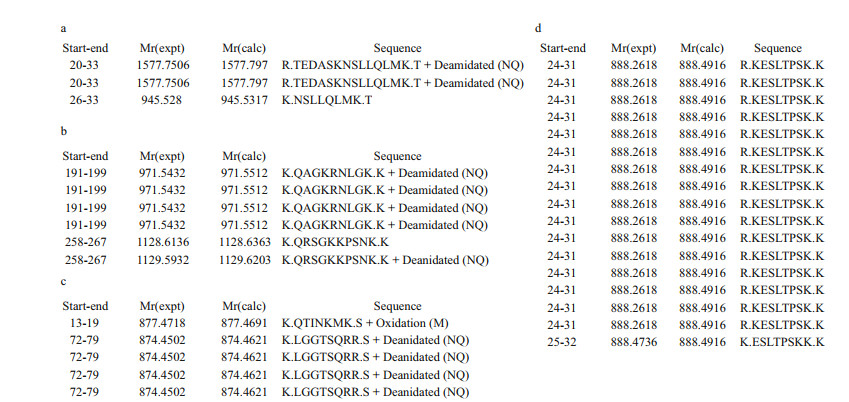
3 RESULT 3.1 Isolation and identification of G. platifrons proteins binding with LPSLPS pull-down assay was conducted to isolate G. platifrons proteins with LPS-binding activity in gills. As a result, massive proteins with molecular weight ranging from 15–20 kDa and 40–75 kDa were obtained in the LPS pull-down elution. A total of 208 host gill proteins were then identified by LCMS/MS (Supplementary Table S1). Among them, five immune genes including LRR-1 (ID evm.TU. Super-Scaffold_12.16), TLR2 (ID evm.TU.SuperScaffold_2293.4), integrin (ID evm.TU.SuperScaffold_1642.4), VSP (ID evm.TU.scaffold307_ size820992_obj.18) and matrix metalloproteinase 1 (MMP1, ID evm.TU.Super-Scaffold_1699.3) were identified from LPS elution and urea elution. In detail, 3 fragments of TLR2, 7 fragments of VSP, 5 fragments of MMP1, and 18 fragments of integrin were also identified in LC-MS/MS (Fig. 1).
|
| Fig.1 The eluted proteins were isolated by LPS pull-down assay and identified by LC-MS/MS a. 3 fragments of TLR2; b. 7 fragments of VSP; c. 5 fragments of MMP1; d. 18 fragments of integrin and 10 fragments of LRR-1 that have been published by Chen et al. (2019) were annotated. |
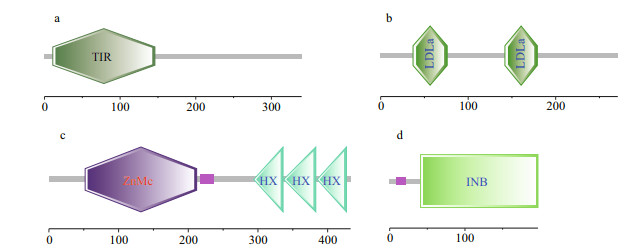
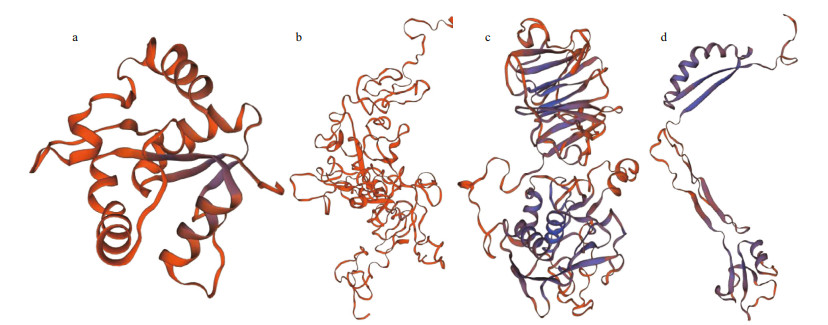
Among the LPS binding genes described above, the structure characteristics and function of LRR-1 have been verified by Chen et al. (2019). The protein structures of TLR2, integrin, MMP1 and VSP proteins were herein analyzed. For TLR2, the full length of its coding region was 1 023 bp and the open reading frame encoded a polypeptide of 340 amino acids residues with molecular weight approximately 38.75 kDa (Supplementary Table S2). The protein sequence had a typical tollinterleukin 1 receptor (TIR) domain but lacked LRR tandem repeats (Fig. 2a). At the same time, the TLR2 protein exhibited a three-dimensional structure with a total of six α-helix and five β-sheets (Fig. 3a). For VSP, the full length of coding region was 816 bp and the open reading frame encoded a polypeptide of 271 amino acids residues with molecular weight approximately 31.40 kDa (Supplementary Table S1). Two low-density lipoprotein receptor class A (LDLa) domains were annotated (Fig. 2b) and a signal peptide (Sec/SPI) was predicted in the VSP protein sequence. At the same time, the VSP protein exhibited a three-dimensional structure with two β-sheets (Fig. 3b). For MMP1, the full length of coding region was 1 260 bp, and it could encode a polypeptide of 420 amino acids residues with molecular weight 48.25 kDa (Supplementary Table S1). A typical ZnMc domain and three tandem hemopexin-like repeat (HX) domains were annotated (Fig. 2c). In addition, a total of five α-helix and 22 β-sheets were predicted in its three-dimensional structure (Fig. 3c). For integrin, the full length of coding region was 594 bp, which can encode a polypeptide of 197 amino acids residues with molecular weight 22.24 kDa (Supplementary Table S1). The protein sequence contained an integrin β subunit domain (Fig. 2d), and exhibited a threedimensional structure with a total of four α-helix and eight β-sheets (Fig. 3d).
|
| Fig.2 Protein domains of PRRs were annotated by SMART a. the protein sequence of TLR2 encoded a typical TIR domain, in consistent with other TLRs; b. the protein sequence of VSP encoded a total of two LDLa domains and a signal peptide; c. one typical ZnMc domain and three tandem HX domains were observed from the MMP1 protein sequence; d. the protein sequence of integrin encoded an integrin β subunit domain. |
|
| Fig.3 Tertiary structures of PRRs were built by Swiss-Model a. a total of six α-helix and five β-sheets were predicted in the tertiary structure of TLR2; b. two β-sheets were predicted in the tertiary structure of VSP; c. a total of five α-helix and 22 β-sheets were predicted in the tertiary structure of MMP1; d. a total of four α-helix and eight β-sheets were predicted in the tertiary structure of integrin. |
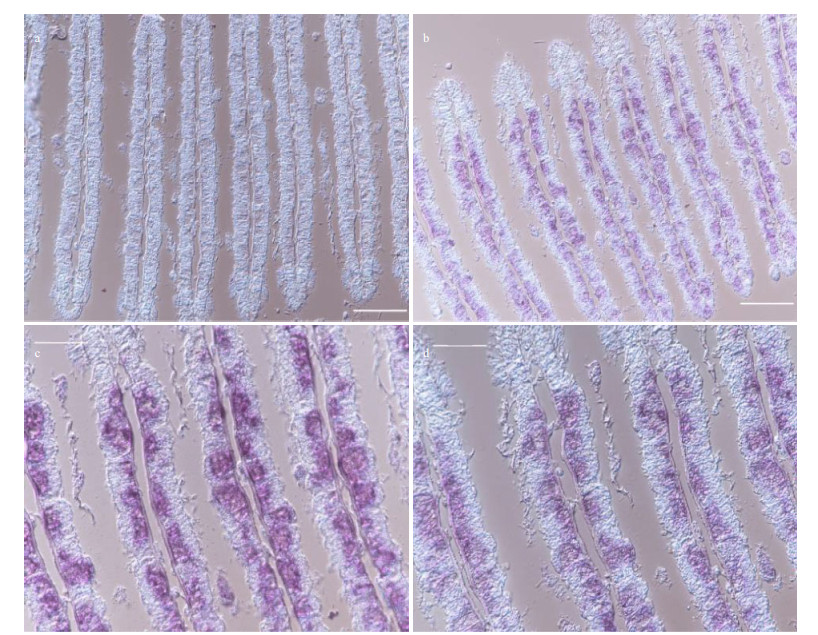
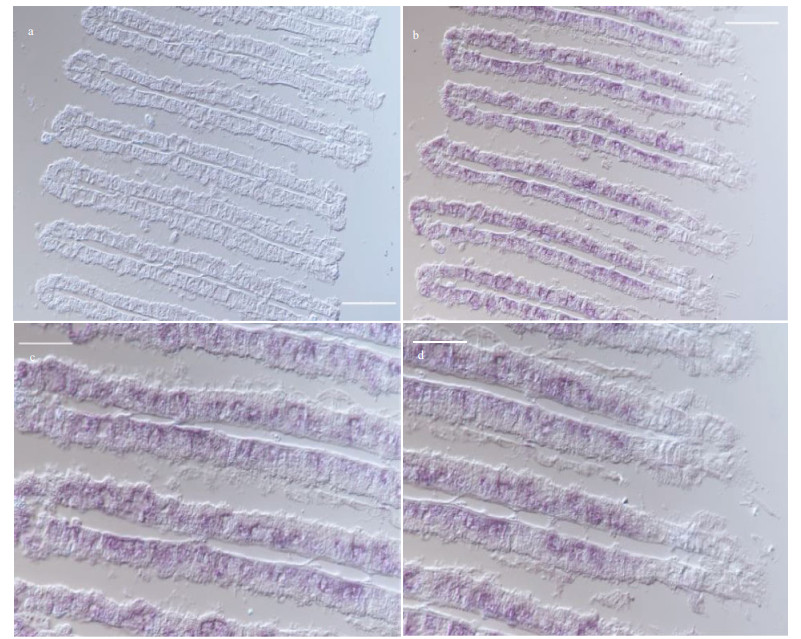
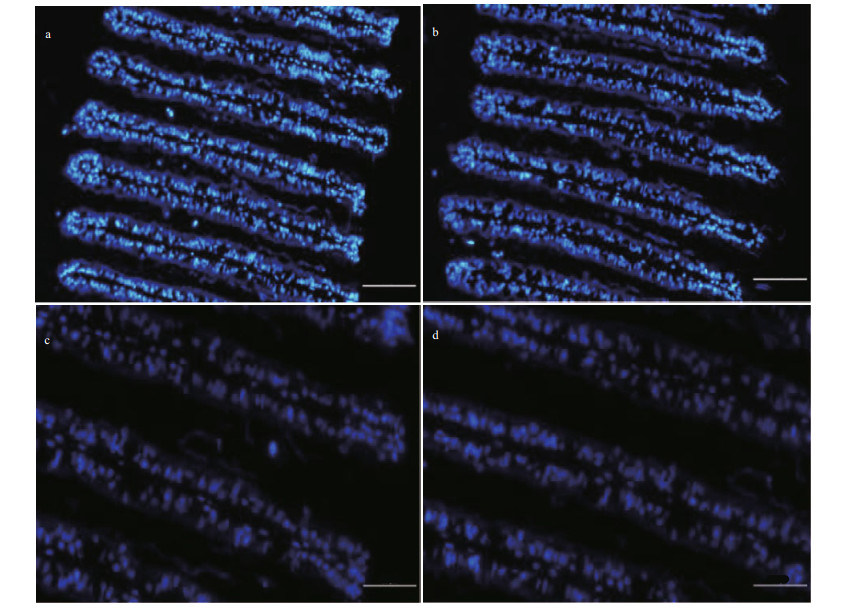
To further investigate the potential role of the above gene products in the symbiosis between hosts and MOBs, the expression patterns of LRR-1, TLR2, and VSP were analyzed by ISH assay in the gill tissues. As a result, mRNA of LRR-1, TLR2, and VSP was exclusively detected in bacteriocytes of gills where the symbionts occurred instead of ciliated cells (Figs. 4–6). What's more, combined with the localization of nuclei in gill cells by DAPI staining, majority of ISH signals were observed near the nuclei of the bacteriocytes (Fig. 7).
|
| Fig.4 ISH of VSP in the gills of G. platifrons a. ISH target to VSP sense mRNA (negative control); b. ISH target to VSP antisense mRNA (positive signal); c & d. the partial enlargements of (b). Scale bar: a–b. 100 μm; c–d. 50 μm. |
|
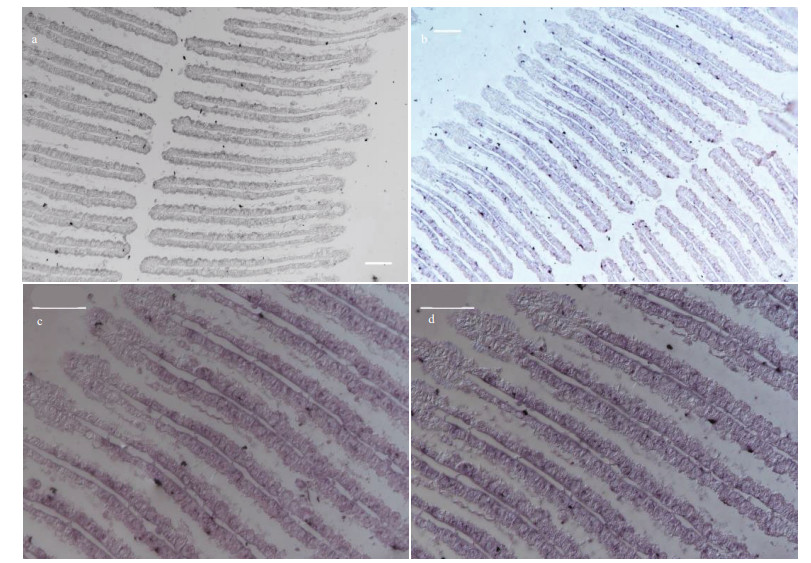
| Fig.5 ISH of LRR-1 in the gills of G. platifrons a. ISH target to LRR-1 sense mRNA (negative control); b. ISH target to LRR-1 antisense mRNA (positive signal); c & d. the partial enlargements of (b). Scale bar: a–b. 100 μm; c–d. 50 μm. |
|
| Fig.6 ISH of TLR2 in the gills of G. platifrons a. ISH target to TLR2 sense mRNA (negative control); b. ISH target to TLR2 antisense mRNA (positive signal); c & d. the partial enlargements of (b). Scale bar: 100 μm. |
|
| Fig.7 FISH with DAPI, showing the distribution of nuclei of gill cells a & b. the different regions of the same slide; c & d. the partial enlargements of (b). Scale bar: a–b. 100 μm; c–d. 50 μm. |
Regardless of massive efforts made in recent years, the lack of cultivable symbiotic MOBs has greatly hindered further research into the molecular mechanisms of the interaction between Bathymodioline and MOBs. Pull-down assay has been proven as an efficient method for verifying molecules involved in pathogen or symbionts recognition (Wu et al., 2011; Li et al., 2016). Given that symbiotic MOBs are γ-proteobacteria with LPS (constituted by lipid A, core oligosaccharide and O polysaccharide) in the outer membrane, LPS pull down assay was herein conducted to capture gill proteins that potentially bound with MOBs. Consequently, 208 proteins ranging from 15–75 kDa were identified, among which five immune genes were also identified demonstrating the reliability of the method. What's more, further verification showed that LRR-1 was likely to play crucial roles in the immune recognition of MOBs (Chen et al., 2019), suggesting the credibility and applicability of the method in Bathymodioline research.
4.2 Immune recognition of symbiontsAs suggested, PRRs mediated immune recognition of symbionts could be the first step in initializing symbiosis. To date, multiple PRRs including Toll-like receptors (TLRs), immunoglobulin superfamily members (IgSF), peptidoglycan recognition proteins (PGRPs), lectins, and complement molecule (C1q) have been reported in Bathymodioline mussels and suggested their irreplaceable role in symbiosis (Bettencourt et al., 2009; Martins et al., 2014; Détrée et al., 2017; Zheng et al., 2017). What's more, it was reported that G. platifrons can maintain a specific symbiotic relationship with MOBs. However, invertebrates are in lack of acquired immunity or immunoglobulins (Cerenius and Söderhäll, 2013). It was therefore speculated G. platifrons might recognize its symbionts through the arrangement and combination of multiple PRRs, which is worthy further investigation. However, none of them was further verified either in vitro or in vivo. Two candidate PRRs including LRR-1 that has been verified by Chen et al. (2019) and TLR2 in present study were isolated by LPS pull-down assay and further characterized by analyzing their structures and performing ISH.
LRR-1 is a classic member of leucine-rich repeat family with diverse functions in host immunity. As found by Chen et al. (2019), LRR-1 might serve as an intracellular recognition receptor for the symbionts by binding with LPS structure. Here, the distribution of LRR-1 was further confirmed by ISH. In consistent with previous conclusions, ISH results showed that LRR-1 was expressed exclusively in bacteriocytes, further demonstrating that LRR-1 could play an important role in immune recognition symbiotic MOBs.
Besides, TLR2 is known as a typical member of TLR family that locates on the cell membrane, and could play important roles in the either pathogen recognition or symbionts recognition by binding with microbe-associated molecular patterns (Akira and Takeda, 2004). Genomic and transcriptome information has showed that TLR family, including TLR2 members, was highly expanded or expressed in G. platifrons and might be crucial in immune recognition of symbionts (Wong et al., 2015; Sun et al., 2017). In consistency, TLR2 was herein found exclusively expressed in bacteriocytes with significant binding activity of LPS. As reported, TLR usually consists of an extracellular LRR domain and an intracellular TIR domain, and can interact with MyD88 to activate downstream intracellular signaling pathways (Kawai and Akira, 2010). In the present study, G. platifrons TLR2 was only found encoding a typical TIR domain which is believed to further trigger downstream signals in bacteriocytes and activates symbiosis related immune response.
4.3 Phagocytosis of symbiontsAfter host immune recognition of symbionts, phagocytosis or endocytosis of symbionts was suggested to be initiated. In our study, two immune genes involving in phagocytosis processes including integrin and VSP were identified by pull-down assay. Integrins are transmembrane receptors that facilitate cell-extracellular matrix (ECM) adhesion. Upon ligand binding, integrins can activate signal transduction pathway that initiate cell phagocytosis (Giancotti and Ruoslahti, 1999). Integrins usually consist of two subunits, α subunit, and β subunit. It was suggested that α subunit could bind with divalent cations that regulate integrin activity. Meanwhile, β subunit usually has four cysteine-rich repeats and that could promote cell adhesion or cell-cell interactions (Humphries, 2000). The integrin protein isolated by LPS pull-down was found containing a typical β subunit. It is therefore speculated that the integrin protein might activate the phagocytosis of symbiotic MOBs through binding with LPS structure.
VSP is known to be involved in the intracellular sorting and delivery of soluble vacuolar proteins, which is essential for late endosome and lysosome assembly and for numerous endolysosomal trafficking pathway (Kolesnikova et al., 2009). In our study, VSP protein, with two LDLa domains and a signal peptide, was also identified in LPS pull-down assay. Though the binding site of VSP against LPS remained unknown, ISH assay demonstrated unique expression pattern of VSP in bacteriocytes, implying its participation in the phagocytosis and transport of symbiotic MOBs in bacteriocytes.
4.4 Digestion of symbiontsAs demonstrated, the Bathymodioline mussels could obtain majority of nutrient needed from their endosymbionts via either "farming" or "milking" way (Fiala-Médioni et al., 2002; Kádár et al., 2008). Though the exact way remains largely unknown, lysosome mediated protein digestion is irreplaceable. Here, MMP1, a key proteinase involving in digestion of proteins, was identified by LPS pull-down assay. MMPs are calcium-dependent zinc-containing endopeptidase, which have been widely identified in vertebrates, invertebrates and plants (Verma and Hansch, 2007). These enzymes can directly digest various extracellular matrix proteins and other bioactive molecules, and play essential roles in cell differentiation and motility, as well as in inflammation and wound healing (Van Lint and Libert, 2007). The MMPs usually contain three conserved domains including pro-peptide domain, catalytic domain and hemopexin-like C-terminal domain. The MMP molecule isolated by LPS pull-down was identified as MMP1, which has a typical ZnMc domain and three tandem HX domains. Given the crucial role of MMPs in the digestion of extracellular matrix in multiple processes, it is inferred that MMP1 may be responsible for the degradation of symbionts directly or indirectly and further promoting the nutrient transferring into G. platifrons cells.
5 CONCLUSIONIn this study, through LPS pull-down assay, we identified four new candidate immune genes that may participate in the symbiosis between G. platifrons and its symbiotic MOBs in addition to LRR-1: VSP, TLR2, MMP1, and integrin. ISH assay was further conducted to investigate the expression patterns of LRR-1, TLR2, and VSP. Consequently, it was found that LRR-1, TLR2 and VSP were expressed exclusively in gill bacteriocytes where symbionts are harbored. Meanwhile, host immune recognition, phagocytosis and lysosome-mediated digestion were suggested to be modulated by above genes. Collectively, the present study has proposed an effective method for screening symbiosis-related immune genes in G. platifrons considering the lack of culturable symbiotic MOBs. Moreover, results obtained herein have also shed new insights into the biological process of symbiosis between G. platifrons and symbiotic MOBs.
6 DATA AVAILABILITY STATEMENTAll data generated and/or analyzed during the study are available from the corresponding author on reasonable request.
7 ACKNOWLEDGMENTWe thank all the crews onboard the R/V Kexue for their assistance in sample collection and all the laboratory staff for continuous technical advice and helpful discussions.
Akira S, Takeda K. 2004. Toll-like receptor signalling. Nature Reviews Immunology, 4(7): 499-511.
DOI:10.1038/nri1391 |
Barry J P, Buck K R, Kochevar R K, Nelson D C, Fujiwara Y, Goffredi S K, Hashimoto J. 2002. Methane-based symbiosis in a mussel, Bathymodiolus platifrons, from cold seeps in Sagami Bay, Japan. Invertebrate Biology, 121(1): 47-54.
DOI:10.1111/j.1744-7410.2002.tb00128.x |
Bettencourt R, Dando P, Collins P, Costa V, Allam B, Serrão Santos R. 2009. Innate immunity in the deep sea hydrothermal vent mussel Bathymodiolus azoricus. Comparative Biochemistry and Physiology Part A:Molecular & Integrative Physiology, 152(2): 278-289.
DOI:10.1016/j.cbpa.2008.10.022 |
Bettencourt R, Pinheiro M, Egas C, Gomes P, Afonso M, Shank T, Serrão Santos R. 2010. High-throughput sequencing and analysis of the gill tissue transcriptome from the deep-sea hydrothermal vent mussel Bathymodiolus azoricus. BMC Genomics, 11(1): 559.
DOI:10.1186/1471-2164-11-559 |
Brooks J M, Kennicutt Ⅱ M C, Fisher C R, Macko S A, Cole K, Childress J J, Bidigare R R, Vetter R D. 1987. Deep-sea hydrocarbon seep communities: evidence for energy and nutritional carbon sources. Science, 238(4830): 1 138-1 142.
DOI:10.1126/science.238.4830.1138 |
Cerenius L, Söderhäll K. 2013. Variable immune molecules in invertebrates. Journal of Experimental Biology, 216(23): 4 313-4 319.
DOI:10.1242/jeb.085191 |
Chen H, Wang M X, Zhang H, Wang H, Lv Z, Zhou L, Zhong Z S, Lian C, Cao L, Li C L. 2019. An LRR-domain containing protein identified in Bathymodiolus platifrons serves as intracellular recognition receptor for the endosymbiotic methane-oxidation bacteria. Fish & Shellfish Immunology, 93: 354-360.
DOI:10.1016/j.fsi.2019.07.032 |
Chow J, Lee S M, Shen Y, Khosravi A, Mazmanian S K. 2010. Host-bacterial symbiosis in health and disease. Advances in Immunology, 107: 243-274.
DOI:10.1016/B978-0-12-381300-8.00008-3 |
Chu H, Mazmanian S K. 2013. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nature Immunology, 14(7): 668-675.
DOI:10.1038/ni.2635 |
Corliss J B, Dymond J, Gordon L I, Edmond J M, von Herzen R P, Ballard R D, Green K, Williams D, Bainbridge A, Crane K, van Andel T H. 1979. Submarine thermal springs on the Galápagos Rift. Science, 203(4385): 1 073-1 083.
DOI:10.1126/science.203.4385.1073 |
Danovaro R, Corinaldesi C, Dell'Anno A, Snelgrove P V R. 2017. The deep-sea under global change. Current Biology, 27(11): R461-R465.
DOI:10.1016/j.cub.2017.02.046 |
Danovaro R, Snelgrove P V R, Tyler P. 2014. Challenging the paradigms of deep-sea ecology. Trends in Ecology & Evolution, 29(8): 465-475.
DOI:10.1016/j.tree.2014.06.002 |
Détrée C, Lallier F H, Tanguy A, Mary J. 2017. Identification and gene expression of multiple peptidoglycan recognition proteins (PGRPs) in the deep-sea mussel Bathymodiolus azoricus, involvement in symbiosis?. Comparative Biochemistry and Physiology Part B: Biochemistry and Molecular Biology, 207: 1-8.
DOI:10.1016/j.cbpb.2017.02.002 |
Dubilier N, Bergin C, Lott C. 2008. Symbiotic diversity in marine animals: the art of harnessing chemosynthesis. Nature Reviews Microbiology, 6(10): 725-740.
DOI:10.1038/nrmicro1992 |
Feng D, Cheng M, Kiel S, Qiu J W, Yang Q H, Zhou H Y, Peng Y B, Chen D F. 2015. Using Bathymodiolus tissue stable carbon, nitrogen and sulfur isotopes to infer biogeochemical process at a cold seep in the South China Sea. Deep Sea Research Part I: Oceanographic Research Papers, 104: 52-59.
DOI:10.1016/j.dsr.2015.06.011 |
Fiala-Médioni A, McKiness Z, Dando P, Boulegue J, Mariotti A, Alayse-Danet A, Robinson J, Cavanaugh C. 2002. Ultrastructural, biochemical, and immunological characterization of two populations of the mytilid mussel Bathymodiolus azoricus from the Mid-Atlantic Ridge:evidence for a dual symbiosis. Marine Biology, 141(6): 1 035-1 043.
DOI:10.1007/s00227-002-0903-9 |
Fontanez K M, Cavanaugh C M. 2014. Evidence for horizontal transmission from multilocus phylogeny of deep-sea mussel (Mytilidae) symbionts. Environmental Microbiology, 16(12): 3 608-3 621.
DOI:10.1111/1462-2920.12379 |
Fujiwara Y, Takai K, Uematsu K, Tsuchida S, Hunt J C, Hashimoto J. 2000. Phylogenetic characterization of endosymbionts in three hydrothermal vent mussels:influence on host distributions. Marine Ecology Progress Series, 208: 147-155.
DOI:10.3354/meps208147 |
Giancotti F G, Ruoslahti E. 1999. Integrin signaling. Science, 285(5430): 1 028-1 033.
DOI:10.1126/science.285.5430.1028 |
Halary S, Riou V, Gaill F, Boudier T, Duperron S. 2008. 3D FISH for the quantification of methane- and sulphuroxidizing endosymbionts in bacteriocytes of the hydrothermal vent mussel Bathymodiolus azoricus. The ISME Journal, 2(3): 284-292.
DOI:10.1038/ismej.2008.3 |
Humphries M J. 2000. Integrin structure. Biochemical Society Transactions, 28(4): 311-39.
DOI:10.1042/bst0280311 |
Kádár E, Davis S A, Lobo-da-Cunha A. 2008. Cytoenzymatic investigation of intracellular digestion in the symbiontbearing hydrothermal bivalve Bathymodiolus azoricus. Marine Biology, 153(5): 995-1 004.
DOI:10.1007/s00227-007-0872-0 |
Kawai T, Akira S. 2010. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nature Immunology, 11(5): 373-384.
DOI:10.1038/ni.1863 |
Kennicutt Ⅱ M C, Brooks J M, Bidigare R R, Fay R R, Wade T L, McDonald T J. 1985. Vent-type taxa in a hydrocarbon seep region on the Louisiana slope. Nature, 317(6035): 315-353.
DOI:10.1038/317351a0 |
Kolesnikova L, Strecker T, Morita E, Zielecki F, Mittler E, Crump C, Becker S. 2009. Vacuolar protein sorting pathway contributes to the release of Marburg virus. Journal of Virology, 83(5): 2 327-2 337.
DOI:10.1128/JVI.02184-08 |
Li H, Huang X Y, Zeng Z H, Peng X X, Peng B. 2016. Identification of the interactome between fish plasma proteins and Edwardsiella tarda reveals tissue-specific strategies against bacterial infection. The International Journal of Biochemistry & Cell Biology, 78: 260-267.
DOI:10.1016/j.biocel.2016.07.021 |
Martins E, Figueras A, Novoa B, Serrão Santos R, Moreira R, Bettencourt R. 2014. Comparative study of immune responses in the deep-sea hydrothermal vent mussel Bathymodiolus azoricus and the shallow-water mussel Mytilus galloprovincialis challenged with Vibrio bacteria. Fish & Shellfish Immunology, 40(2): 485-499.
DOI:10.1016/j.fsi.2014.07.018 |
Ponnudurai R, Kleiner M, Sayavedra L, Petersen J M, Moche M, Otto A, Becher D, Takeuchi T, Satoh N, Dubilier N, Schweder T, Markert S. 2017. Metabolic and physiological interdependencies in the Bathymodiolus azoricus symbiosis. The ISME Journal, 11(2): 463-477.
DOI:10.1038/ismej.2016.124 |
Sayavedra L, Kleiner M, Ponnudurai R, Wetzel S, Pelletier E, Barbe V, Satoh N, Shoguchi E, Fink D, Breusing C, Reusch T B H, Rosenstiel P, Schilhabe M B, Becher D, Schweder T, Markert S, Dubilier N, Petersen J M. 2015. Abundant toxin-related genes in the genomes of beneficial symbionts from deep-sea hydrothermal vent mussels. eLife, 4: e07966.
DOI:10.7554/eLife.07966.001 |
Sibuet M, Olu K. 1998. Biogeography, biodiversity and fluid dependence of deep-sea cold-seep communities at active and passive margins. Deep Sea Research Part Ⅱ: Topical Studies in Oceanography, 45(1-3): 517-567.
DOI:10.1016/s0967-0645(97)00074-x |
Sievert S M, Vetriani C. 2015. Chemoautotrophy at deep-sea vents: past, present, and future. Oceanography, 25(1): 218-233.
DOI:10.5670/oceanog.2012.21 |
Sun J, Zhang Y, Xu T, Zhang Y, Mu H W, Zhang Y J, Lan Y, Fields C J, Hui J H L, Zhang W P, Li R S, Nong W Y, Cheung F K M, Qiu J W, Qian P Y. 2017. Adaptation to deep-sea chemosynthetic environments as revealed by mussel genomes. Nature Ecology & Evolution, 1(5): 0121.
DOI:10.1038/s41559-017-0121 |
Van Dover C L, German C R, Speer K G, Parson L M, Vrijenhoek R C. 2002. Evolution and biogeography of deep-sea vent and seep invertebrates. Science, 295(5558): 1 253-1 257.
DOI:10.1126/science.1067361 |
Van Lint P, Libert C. 2007. Chemokine and cytokine processing by matrix metalloproteinases and its effect on leukocyte migration and inflammation. Journal of Leukocyte Biology, 82(6): 1 375-1 381.
DOI:10.1189/jlb.0607338 |
Verma R P, Hansch C. 2007. Matrix metalloproteinases(MMPs): chemical-biological functions and (Q)SARs. Bioorganic & Medicinal Chemistry, 15(6): 2 223-2 268.
DOI:10.1016/j.bmc.2007.01.011 |
Wentrup C, Wendeberg A, Huang J Y, Borowski C, Dubilier N. 2013. Shift from widespread symbiont infection of host tissues to specific colonization of gills in juvenile deepsea mussels. The ISME Journal, 7(6): 1 244-1 247.
DOI:10.1038/ismej.2013.5 |
Won Y J, Hallam S J, O'Mullan G D, Pan I L, Buck K R, Vrijenhoek R C. 2003. Environmental acquisition of thiotrophic endosymbionts by deep-sea mussels of the genus Bathymodiolus. Applied and Environmental Microbiology, 69(11): 6 785-6 792.
DOI:10.1128/aem.69.11.6785-6792.2003 |
Wong Y H, Sun J, He L S, Chen L G, Qiu J W, Qian P Y. 2015. High-throughput transcriptome sequencing of the cold seep mussel Bathymodiolus platifrons. Scientific Reports, 5(1): 16597.
DOI:10.1038/srep16597 |
Wu C L, Söderhäll K, Söderhäll I. 2011. Two novel ficolin-like proteins act as pattern recognition receptors for invading pathogens in the freshwater crayfish Pacifastacus leniusculus. Proteomics, 11(11): 2 249-2 264.
DOI:10.1002/pmic.201000728 |
Xu J C, Jiang S, Li Y Q, Li M J, Cheng Q, Zhao D P, Yang B, Jia Z H, Wang L L, Song L S. 2016. Caspase-3 serves as an intracellular immune receptor specific for lipopolysaccharide in oyster Crassostrea gigas. Developmental & Comparative Immunology, 61: 1-12.
DOI:10.1016/j.dci.2016.03.015 |
Zheng P, Wang M X, Li C L, Sun X Q, Wang X C, Sun Y, Sun S. 2017. Insights into deep-sea adaptations and hostsymbiont interactions: a comparative transcriptome study on Bathymodiolus mussels and their coastal relatives. Molecular Ecology, 26(19): 5 133-5 148.
DOI:10.1111/mec.14160 |
 2020, Vol. 38
2020, Vol. 38



