Institute of Oceanology, Chinese Academy of Sciences
Article Information
- QIN Huan, GUO Qingqing, LIU Chenchen, LI Fenglan, ZHANG Hua, CHU Zihan, WANG Jiangxin, LEI Anping
- Occurrence and light response of residual plastid genes in a Euglena gracilis bleached mutant strain OflB2
- Journal of Oceanology and Limnology, 38(6): 1858-1866
- http://dx.doi.org/10.1007/s00343-019-9159-9
Article History
- Received Jun. 18, 2019
- accepted in principle Sep. 23, 2019
- accepted for publication Nov. 4, 2019
2 Basis International School SZ, Shenzhen 518000, China
Euglena is a unicellular protist that shows characteristics of both plants and animals (Bodyl, 1996). Most Euglena species contain fully developed chloroplasts when autotrophically grown through the photosynthesis under light, and they can also heterotrophically grow by absorbing organic matters in the dark (Tucci et al., 2010). The chloroplasts of Euglena are believed to be originated from a unicellular green alga endosymbiosis (Sulli et al., 1999; Ahmadinejad et al., 2007; Yoshida et al., 2016; Zakryś et al., 2017). Unlike chloroplasts of other algae and higher plants, the chloroplasts of Euglena have poor stability and are easy to lose under stresses, such as antibiotics, heat, UV, etc., but retain parts of the plastid genome (Kivic and Vesk, 1974; Heizmann et al., 1981; Wang et al., 2002, 2004). Euglena cells that lost a part of their photosynthesis-related genes become bleached mutants and can stably grow and multiply (Gockel et al., 2000). Therefore, Euglena is an ideal model species for studying the function, development, and endosymbiosis of chloroplast. Euglena gracilis is the most popular and typical species, and it is usually chosen as a model for Euglena.
When Euglena grows in the dark, its chloroplasts are poorly developed. Once returned to light, Euglena turns green, reestablishes its chloroplasts, and begins to function again as a photosynthetic organism (Kivic and Vesk, 1972). In addition, when treated with UV or ofloxacin (Ofl), E. gracilis cells will permanently lose their chloroplasts' photosynthetic functionality and become bleached mutants, while most mutants will still have residual plastids and partial plastid genomes (Sulli et al., 1999; Ahmadinejad et al., 2007; Yoshida et al., 2016; Zakryś et al., 2017). Residual plastids were observed in a bleached mutant of E. gracilis by transmission electron microscopy (Kivic and Vesk, 1974; Wang et al., 2004), and a deleted chloroplast genome was further demonstrated by Heizmann's hybridization method (Heizmann et al., 1976, 1982; Hussein et al., 1982). Moreover, chloroplast rRNA was detected in 25 bleached mutants, and the majority of these rRNAs are similar in number to those found in wild type E. gracilis (Heizmann et al., 1976, 1982; Hussein et al., 1982).
Euglena gracilis plastid genome consists of 143 170 bp, a total of 97 proteins and gene loci, including 46 protein-coding genes, and 51 RNA-coding genes (Bennett and Triemer, 2015; Hadariová et al., 2017). Except for the presence of three copies of the rRNA gene, each gene is present in a single copy matter (Hallick et al., 1993; Hadariová et al., 2017). Euglena longa is a naturally-occurring, colorless, non-photosynthetic euglenoid, which looks like bleached E. gracilis in appearance. The discovery of a circular 73-kb plastid genome in E. longa, which is about half the size of the circular 143 kb in E. gracilis, suggested that the colorless E. longa could not simply be a naturally bleached E. gracilis (Gockel et al., 1994; Eberhard et al., 2002). All the genes encoding photosynthetic proteins were lost from the E. longa plastid genome except for the rbcL gene encoding the large subunit of ribulose-1, 5-bisphosphate carboxylase/oxygenase (Gockel et al., 1994). Furthermore, the divergence between the plastid genomes in E. gracilis, and E. longa appears to have involved in a selective loss and rearrangement of E. longa plastid genes rather than an expected random loss (Siemeister and Hachtel, 1989; Gockel et al., 1994; Eberhard et al., 2002). It is essential to understand if the loss of plastid genes in the bleached mutant of E. gracilis was selective.
Most previous studies on the bleached mutants of E. gracilis focused on the presence or absence of plastid genomes, and few have studied the plastid genes and their functions (Buetow, 1982; Vesteg et al., 2009). To understand the function of residual plastid DNA, the present study compared the expression level of their genes in WT and bleached mutant under light stimulated conditions.
2 MATERIAL AND METHOD 2.1 Strains and culture conditionsEuglena gracilis CCAP 1224/5Z was purchased from CCAP (Culture Collection of Algae and Protozoa, UK) and maintained in our laboratory. WT and the bleached mutants were grown in EG medium (Supplementary Table S1) under the condition of 12 h/12 h light (100±10 μmol/(m2∙s)/dark at 22℃. Solid plates were supplemented with 2% agar.
2.2 Experimental set-upTo obtain the bleached mutants, Ofl was added to the medium to a final concentration of 100 μmol/L, and 100 cells were spread on the EG solid medium plate. After 7 d, five white mutants (OflB1, OflB2, OflB3, OflB4, and OflB5) were picked from the plate and transferred to liquid EG medium for continuous cultivation. These bleached colonies have been cultivated for more than 2 years in our lab with no reversed phenotype (greening), indicating the phenotypic stability of these mutants, and that the bleaching was permanent.
The growth curves of the WT and five bleached mutants of E. gracilis were monitored. The five mutants were further confirmed bleached by UV-visible spectrophotometer, and their residual plastid genes were determined by PCR. Quantitative real time PCR (qRT-PCR) was used to obtain expression level of the plastid residual genes in WT and one of the bleached mutant (OflB2), which were grown for 5 d under dark conditions and then remained in the dark or stimulated by light conditions for 0.5 and 3 h. Thus, five treatments were labeled as D0, D0.5, D3, L0.5, and L3. All the control and experimental samples were in triplicates.
2.3 Detection of plastid genes with PCRThe total DNA of WT and the five mutants was extracted using the TransGen Biotech DNA isolation Kit (EE141-01). Briefly, cells grown at the log phase were harvested by centrifugation at 8 000×g for 2 min, and resuspended in 400 μL of Plantzol containing 10 μL of Proteinase K and 10 μL of RNase A (10 mg/mL). The suspension was then incubated at 55℃ for 0.5 h to completely lyse the cells. An equal volume of phenol:chloroform:isoamyl alcohol (25:24:1) was added, vortexed and mixed, centrifuged at 12 000×g for 5 min, and the supernatant was removed to a new Eppendorf tube. An equal volume of isopropanol was added, vortexed well and centrifuged at 12 000×g for 15 min at 4℃. The precipitated DNA was washed with 70 % ethanol and resuspended in 50 μL deionized water for further use.
Primers were designed for PCR and qRT-PCR according to the chloroplast genome of E. gracilis (Hallick et al., 1993) and were synthesized by Guangzhou Ige Biotechnology Company (Supplementary Table S2). A total of 50 μL of PCR reaction system was set up: 25 μL of 2×Es Taq MasterMixa (Dye), 1 μL of upstream primer (10 μmol/L), 1 μL of downstream primer (10 μmol/L), 2 μL of 300 ng/μL DNA template, and 21 μL of sterilized ddH2O. The PCR amplification was started with denaturation at 95℃ for 3 min, followed by 35 cycles of 10 s at 95℃, 20 s at 56℃, and 10 s at 72℃, and a final extension of 5 min at 72℃. The PCR products of rpl16-rpl5, rpl5-rps14 and rps8-rps2 from WT and mutants were purified and sequenced by Guangzhou Ige Biotechnology company.
2.4 qRT-PCRqRT-PCR was conducted to obtain expression level of the plastid residual genes of WT and OflB2 bleached mutant: rps2, rps3, rps8, rps14, rpl5, rpl14, psbA, psaC, petB, rbcL, and 5s rRNA. Nuclear Actin gene was used as controls.
Total RNAs of WT and OflB2 mutant were extracted using the TaKaRa RNAiso Plus Kit (D9108A). Briefly, cells were harvested by centrifugation at 8 000×g for 2 min, and resuspended in 1 mL of RNAiso Reagent. 200 μL of chloroform (1/5 volume of RNAiso Reagent) was added, vortexed for 15 s and stood at room temperature for 5 min, centrifuged at 12 000 r/min for 15 min at 4℃, and the supernatant was transferred to another new Eppendorf tube. An equal volume of isopropanol was added, mixed gently and stood at room temperature for 10 min, centrifuged at 12 000×g for 10 min at 4℃, and the supernatant was removed. The precipitated RNA was washed with 75% ethanol, resuspended in 20-μL RNase-free water, and stored at -80℃ if not used immediately. Three microgram RNA of each sample was quantitatively reverse transcribed into cDNA using TaKaRa's PrimeScriptTM RT reagent Kit with gDNA eraser (RR047A). The cDNA of each sample was used for Quantitative PCR (qPCR). Actin gene from E. gracilis was used as an internal control for the data normalization. qPCR reaction system was set up: 5-μL Fast SYBRTM Green Master Mix (2x), 0.8-μL upstream primer (10 μmol/L), 0.8-μL downstream primer (10 μmol/L), 3-μL template, and 0.4 μL of sterilized ddH2O. The profile of qPCR included 40 cycles of 95℃ for 15 s and 60℃ for 30 s. Each qRT-PCR measurement was performed in triplicates. Expression analysis was performed by the 2-ΔΔCt method using the analysis software provided by the Bio-Rad qPCR instrument.
2.5 Statistical analysisThe data of light/dark and different time treatments of WT and OflB2 mutant were analyzed using a parametric two-way analysis of variance (ANOVA) (light/dark treatment n=2, time treatment n=3). To compare all differences among three times during light and dark treatment, one-way analysis of variance (ANOVA) was used. The statistical analysis was carried out by SPSS 17.0 for windows.
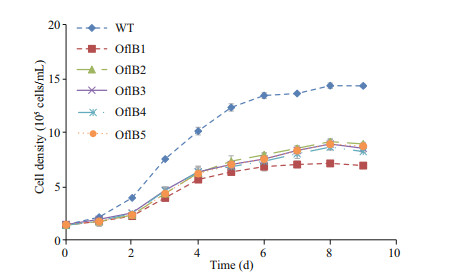
3 RESULT 3.1 Characteristics of the bleached mutants of E. gracilisAll the five mutants (OflB1, OflB2, OflB3, OflB4 and OflB5) presented similar growth curves but their growths were obviously lower than that of WT (Fig. 1). All samples were inoculated at the same initial cell density of 1.4×105 cells/mL, but since the mutants seemed to experience a more obvious lagged growth before entering into the logarithmic growth phase, the cell density of WT was much higher than that of the mutants from day 2 to 8 (Fig. 1). The highest cell density of WT (1.36×106 cells/mL) was achieved at the end of the logarithmic phase at day 8 and was much higher than that of the mutants, which was only 0.9×106 cells/mL for the OflB2 mutant (Fig. 1).
|
| Fig.1 Growth curves of WT and the five mutants (OflB1–5) |
UV-visible spectrophotometer was used to scan the pigment absorption spectrum of WT and the five mutants. Results showed that WT had high absorption in the blue and red regions (sharp absorption peaks at 433 nm and 665 nm) (Supplementary Fig.S1), which coincided with the absorption spectrum of intact chloroplast that contains predominantly chlorophyll and some carotenoids. However, no absorption peak spectrum was observed in all five mutants.
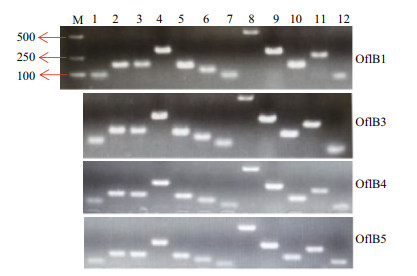
3.2 Detection of plastid genes with PCRConsidering the bleached phenotype of these mutants, it would be interesting to examine whether and how many genes were preserved in the five mutants. We first examined 50 chloroplast-relevant genes (Supplementary Table S2) in a randomly-selected mutant OflB2 using PCR, and 13 genes (rps2, rpl5, rps14, petB, psbA, 5s rRNA, rpl14, rps3, psaC, rpl16, rps8, actin, and rbcL) (Supplementary Table S2) were detected (Fig. 2). Among these genes, rps2, rps3, rps8, rps14, rpl5, rpl14 and rpl16 encode for ribosomal proteins, psaC and psbA for photosynthetic proteins, petB for b6 protein and the core component of the cytochrome b6f protein complex, and rbcL for the large subunit of RuBisCO. We did not test the other 37 genes in all the other 4 mutants, while we focused on the 13 genes existed in OflB2. So we then examined these 13 genes in the other four mutants (OflB1, OflB3, OflB4, and OflB5), and the results indicated the existence of these genes in all mutants (Fig. 3).

|
| Fig.2 Detection of residual plastid genes in OflB2 mutant M: DNA marker; 1-12: representing 5s rRNA, rps8, rpl5, psaC, petB, rps3, rps2, rps14, psbA, rpl14, rbcL, and rpl16 genes respectively. |
|
| Fig.3 Detection of residual plastid genes in OflB1, OflB3, OflB4, and OflB5 mutants M: DNA marker; 1-12: representing 5s rRNA, rps8, rps3, rps2, rps14, psaC, rpl14, rbcL, rpl5, psbA, petB, rpl16 genes respectively. |
Among the 12 genes detected, 6 genes (rpl16, rpl14, rpl5, rps8, rps14, and rps2) were closely distributed to each other, which made it possible to explore the rpl16-rps2 inter-regions between these 6 genes with appropriate primers (Supplementary Fig. S2). These regions of WT and OflB2 were therefore amplified with the primer pairs of rpl16-rpl5, rpl5-rps14, and rps8-rps2, and the PCR products were sequenced and assembled, indicating that the lengths of rpl16-rps2 region in WT and OflB2 mutant were 4 122 bp and 4 114 bp, respectively (Supplementary Sequence S1). Alignment analysis with BLAST showed that the similarity between the two sequences were 0.99, and 51 mutations with 10 deletions/insertions and 41 point mutations existed in OflB2 mutant compared with WT. Interestingly, most of these mutations existed in the rps8-rps2 region (Supplementary Sequence S1).
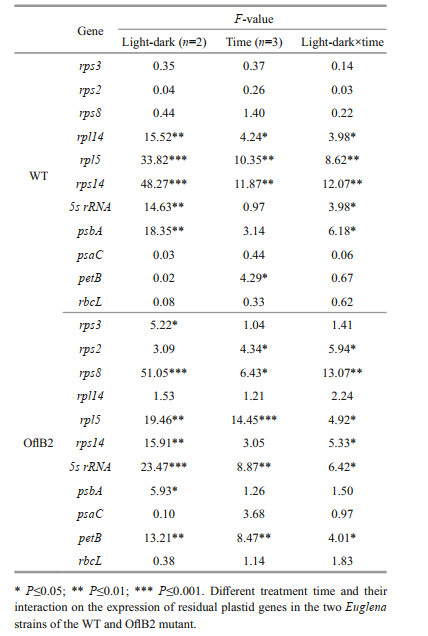
3.3 Expression of residual plastid genesThe transcription levels of 11 plastid genes (rps2, rps3, rps8, rps14, rpl5, rpl14, psbA, psaC, petB, rbcL, and 5s rRNA) (expect rpl16, due to its low expression level) in WT and OflB2 mutant under dark or light-stimulation conditions were analyzed by qRT-PCR. According to the two-way ANOVA test, effects of light-dark and different treatment time on and their interactions with the expression of residual plastid genes were gene specific (Table 1). The light-dark effect in WT was only significant on five genes, which were rpl14, rpl5, rps14, 5s rRNA, and psbA; and the time effects in WT were significant on rpl14, rpl5, rps14, and petB. The light-dark effect in OflB2 was significant on seven genes, including rps3, rps8, rpl5, rps14, 5s rRNA, psbA, and petB, and the time effects in OflB2 were significant on rps2, rps8, rpl5, 5s rRNA, and petB. The light-dark or time showed insignificant effects on the other genes.
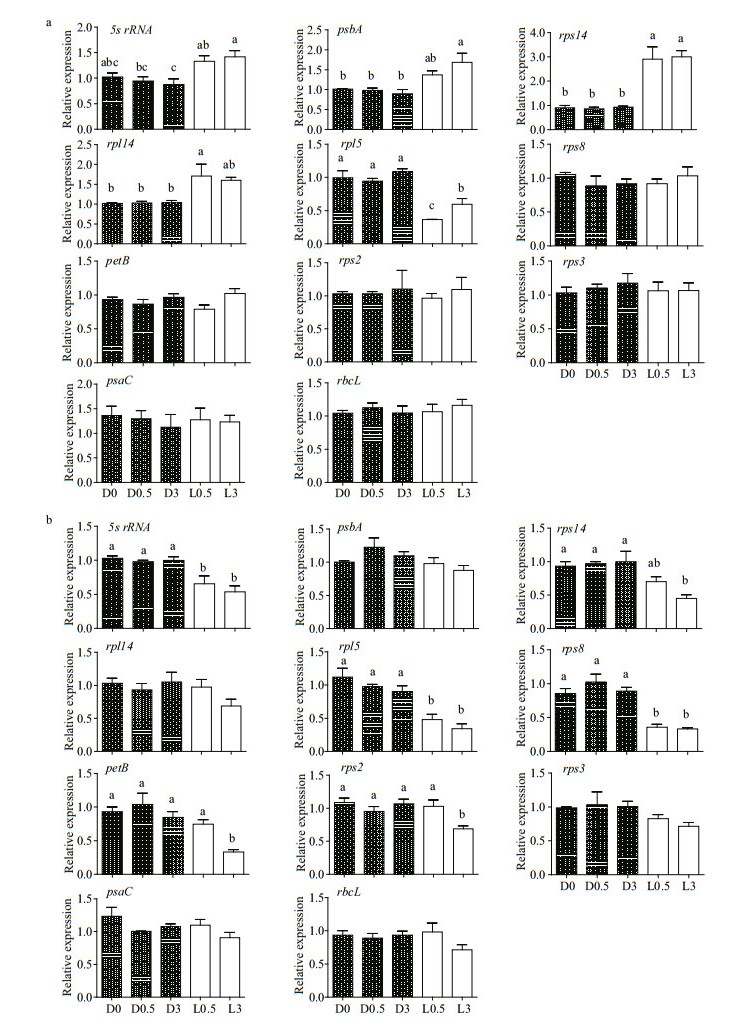
Results of multiple comparisons showed that the expression of all the 11 plastid genes in WT and OflB2 mutant did not change significantly in the dark, and then continued in the dark for 0.5 h and 3 h (Fig. 4a & b) (P > 0.05), indicating that the expression of these genes were stable in dark conditions, whereas response to light stimulation varied among the genes and was different between WT and OflB2 mutant. For WT, only gene rpl5 was significantly down-regulated, whereas four genes (5s rRNA, rps14, rpl14i, and psbA) were significantly up-regulated under light stimulation conditions (Fig. 4a, Table 1) (P≤0.05). For OflB2 mutant, the expression levels of genes 5s rRNA, petB, rpl5, rps8, rps14, and rps2 were significantly down-regulated under light stimulation conditions, and the decrease was more remarkable in longer light stimulation: the expression of genes petB and rps2 remained more or less the same as that in dark conditions after 0.5-h light stimulation but dropped sharply after 3-h light stimulation (Fig. 4b, Table 1) (P≤0.05). No significant difference was observed in the expressions of the other six (rps2, rps3, rps8, psaC, petB, and rbcL) and five (rps3, rpl14, psbA, psaC, and rbcL) genes in WT and OflB2 from dark to light stimulations, respectively (P > 0.05) (Fig. 4a & b).
|
| Fig.4 Expression of residual plastid genes in WT (a) and OflB2 (b) under dark (D) or light (L) conditions for 0.5 and 3 h (mean and standard deviation, n=3) Different letters on the top of the bar indicate the means were significant different at P≤0.05 (one-way ANOVA); no letters on the top of the bar indicate not significant. |
Ofl is considered as a fluoroquinolone antibacterial drug that can cause loss of chloroplast photosynthetic function in Euglena. It is an inhibitor of DNA gyrase (bacterial topoisomerase type II) and directly inhibits plastid DNA replication. As the cells divide, the new Euglena cells lose their chloroplasts and cannot effectively perform photosynthesis, forming a white bleached mutant (Krajčovič et al., 1989; Križková et al., 1998; Schwartzbach and Schiff, 1974). However, Ofl did not bleach two green algal species, Scenedesmus obliquus (Qin et al., 2012) and Chlamydomonas (lab data, unpublished). In addition, the presence of Ofl has little effect on the growth of E. gracilis (Schwartzbach and Schiff, 1974; Krajčovič et al., 1989; Križková et al., 1998), while it has a toxic effect on the growth and physiological status of S. obliquus (Qin et al., 2012). The mechanism by which Ofl can bleach Euglena but other species of algae remains unclear.
The decrease of chlorophyll is an important indicator of the bleaching of Euglena (Thomas and Ortiz, 1995). The absorption peak of WT indicates that the pigment of WT contains chlorophyll and carotene, which is consistent with the study of the pigment composition of the predecessors (Krinsky and Goldsmith, 1960). However, the mutants did not contain any chlorophyll and carotene, which is consistent with our previous observations (Wang et al., 2002, 2004). The growth of the bleached mutants was much slower than that of WT in the light (Fig. 1), as E.gracilis can be both autotrophic and heterotrophic, while bleached mutants have lost their chloroplasts' function and could not perform photosynthesis but only grow heterotrophically. In contrast to our results, the previous research reported that the presence of Ofl did not affect the growth of E. gracilis (Križková et al., 1998; Hadariová et al., 2017) perhaps caused by the heterogeneity of residual plastid genes in different Ofl mutants. We used different growth medium and at different growth conditions from the other researches (Križková et al., 1998; Hadariová et al., 2017). As stated in Section 2.1, our strains were grown in EG medium under the condition of 12 h/12 h light (100±10 μmol/(m2∙s))/dark at 22℃. Križková et al. (1998) used modified CM supplemented with 0.5% sodium acetate at 27℃ under permanent illumination (2 000 lx). Hadariová et al. (2017) used modified CM medium supplemented with 0.8% ethanol at 27℃, with continuous illumination (30 μmol/(m2∙s)) or in the dark. And these may be the reasons attribute to the difference. The effects and mechanism of Ofl on algal growth need to be further elucidated.
Using the known E. gracilis chloroplast genome (Hallick et al., 1993), 50 genes were selected, and PCR was carried out using the DNA of WT and OflB2 mutant as templates. Only 12 genes were detected, and the remaining 38 genes were all detected in WT but not in the OflB2 mutant (data not shown). However, failing to detect these genes in the OflB2 mutant does not indicate that the OflB2 mutant has lost the undetected genes. It is possible that these genes may have been mutated or rearranged in the regions of selected primers. Similarly, the detection of genes in the OflB2 mutant does not mean that the OflB2 mutant contains the entire genetic sequence.
It is well known that some Euglena bleached mutants contain residual plastids and partial plastid genomes (Heizmann et al., 1981; Gockel et al., 2000; Wang et al., 2002, 2004). We found that the E. gracilis mutants contain 12 residual plastid genes, namely rps2, rps3, rps8, rps14, rpl5, rpl14, psbA, psaC, rpl16, petB, rbcL, and 5s rRNA (Figs. 2 & 3). In previous reports, one mutant sm5 retained 16s rRNA, psbD, psaA, rpl16, rps9i, and rpoB genes, while the other mutants only retained 16s rRNA gene and rpoB region but lost genes rpl3, rbcL, and atpE. 16s rRNA gene and rpoB are next to the replication origin, and therefore they were retained in most mutants (Wang et al., 2004). However, in this study, rps3, rpl16 and rbcL genes were retained in all mutants, and 16s rRNA, psbD, psaA, rps9, rpoB, atpE genes were not detected. Moreover, in our experiment, the rps3, rpl16, and rbcL genes were retained in all mutants. The copy number of rrn16, rrn23, rpl2, tufA, psbC, rbcL, and rpoC2 genes was reduced in the presence of Ofl in the E. gracilis, and the copy number of rpl16 gene remained unchanged (Gockel et al., 2000). In addition, WgmZOflL mutant has also retained rpl14 and rpl5 genes which are present downstream from rpl16 gene, while it does not possess 16s rRNA gene (Krnáčová et al., 2015; Oldenburg and Bendich, 2016). This is consistent with our experiments, and our five mutants also retained the rpl16, rpl5, and rpl14 genes. Our residual plastid genes were different from that of previous research, indicating that the residual plastids genes of bleached mutants varied among different treatments, perhaps even in the same treatment different cells may lose different part of the plastid genome. More detailed molecular analyses of stable bleached mutants and after the bleaching of various E. gracilis strains would be needed to explain the phenomenon of differential retention of plastid genes in different mutants.
We planned to amplify the plastid genome of the OflB2 mutant by using long distance PCR with primers derived from the 13 genes detected. However, it was unsuccessful due to the unknown structure of the OflB2 plastid genome and distribution of these genes. Finally, 6 genes (rpl16, rpl14, rpl5, rps8, rps14, and rps2) were found to be closely distributed to each other, which made it possible to explore the rpl16-rps2 region of OflB2 mutant and WT (Supplementary Sequence S1). The DNA sequences of the rpl16-rps2 region of OflB2 mutant and WT were compared by NCBI BLAST. The results showed that the sequence of OflB2 mutant and WT were highly similar with a 1% difference, and 51 mutations with 10 deletions/insertions and 41 point mutations existed in OflB2 mutant compared to WT. The mismatch rate is higher than the error rate of the high-fidelity Taq enzyme (10-6 bp/cycle), and thus the PCR error could be excluded in this study. In addition, divergence of E. longa from E. gracilis may be caused by the gradual excision of small plastid genome fragments under some extreme environments (Bodyl, 1996).
There are 116 genes in the chloroplast genome of E. gracilis; however, the functions of these genes were rarely studied using molecular techniques. Moreover, streptomycin and Ofl inhibit E. longa growth, suggesting that it requires plastid genes to survive, i.e., its plastid is a functional organelle not just a chloroplast remnant (Gockel et al., 2000). In addition, nuclear encoded plastid genes petJ and psbO were not transcribed in an Ofl induced E. gracilis mutant in the light (Vacula et al., 2001). Although substantial regulation of Euglena chloroplast gene expression also occurs at the translational level, the observed patterns in transcriptional levels are possibly significant (Vacula et al., 2001; Idoine et al., 2014; Ebenezer et al., 2017). Our results showed that the expression of rpl5, 5s rRNA, rps14, rpl14, and psbA genes in WT E. gracilis under light-stimulation condition was changed, only rpl5 gene was significantly down-regulated whereas the other four were significantly up-regulated; this indicates that light stimulation leads to an increase of transcription level of some chloroplast genes of E. gracilis, showing significant light responses. Unexpectedly, the expression of rpl5, 5s rRNA, rps2, rps8, and petB genes in the OflB2 mutant were significantly down-regulated under light stimulation conditions, suggesting that light stimulation can cause expression change of photosynthetic genes in the non-photosynthetic OflB2 mutant, perhaps with negative effects. In WT and OflB2 mutant, the same plastid genes responded to light stimulation in different ways. We could not completely remove the residual plastid DNA of bleached mutants after a long-term of 2 years cultivation, proposing that residual plastid DNA is likely to have functions other than photosynthesis, such as synthesis of some certain essential compounds. This is very interesting and the investigation of the OflB2 mutant residual plastid genes' function is thus needed.
5 CONCLUSIONIn summary, OflB2 bleached mutant retains part of the plastid DNA even after long term (2 years) exposure to Ofl, and the plastid genes respond to light at the transcriptional level. Moreover, chloroplast and plastid genes in WT and OflB2 mutant have different responses to light stimulation. Our research will shed light on understanding the functions of residual plastid DNA in non-photosynthetic microorganisms and evolution of the chloroplast in green microalgae.
6 DATA AVAILABILITY STATEMENTAll data generated and/or analyzed during this study are included in this published article and in its supplementary information files.
7 ACKNOWLEDGMENTThe authors gratefully acknowledge the supports from the Instrumental Analysis Center of Shenzhen University (Xili Campus).
8 AUTHOR CONTRIBUTIONWANG Jiangxin contributed to the conception and design of the study. QIN Huan performed the experiments. LEI Anping, GUO Qingqing, LIU Chenchen, and LI Fenglan performed the statistical analysis. QIN Huan wrote the first draft of the manuscript. WANG Jiangxin, QIN Huan, CHU Zihan, ZHANG Hua, and LEI Anping wrote the sections of the manuscript. WANG Jiangxin, CHU Zihan, and LEI Anping contributed to manuscript revision. All authors read and approved the submitted version.
9 CONFLICT OF INTERESTThe authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Electronic supplementary materialSupplementary material (Supplementary Tables S1–S2, Figs.S1–S2, Sequence S1) is available in the online version of this article at https://doi.org/10.1007/s00343-019-9159-9.
Ahmadinejad N, Dagan T, Martin W. 2007. Genome history in the symbiotic hybrid Euglena gracilis. Gene., 402(1-2): 35-39.
DOI:10.1016/j.gene.2007.07.023 |
Bennett M S, Triemer R E. 2015. Chloroplast genome evolution in the Euglenaceae. J. Eukaryot. Microbiol., 62(6): 773-785.
DOI:10.1111/jeu.12235 |
Bodyl A. 1996. Is the origin of Astasia longa an example of the inheritance of acquired characteristics?. Acta Protozool., 35(2): 87-94.
|
Buetow D E. 1982. The Biology of Euglena, Vol. III. Academic Press, New York. p.157-195.
|
Ebenezer T E, Carrington M, Lebert M, Kelly S, Field M C. 2017. Euglena gracilis genome and transcriptome: organelles, nuclear genome assembly strategies and initial features. In: Schwartzbach S D, Shigeoka S eds. Euglena: Biochemistry, Cell and Molecular Biology. Springer, Cham. p.125-140.
|
Eberhard S, Drapier D, Wollman F A. 2002. Searching limiting steps in the expression of chloroplast-encoded proteins: relations between gene copy number, transcription, transcript abundance and translation rate in the chloroplast of Chlamydomonas reinhardtii. Plant J., 31(2): 149-160.
DOI:10.1046/j.1365-313X.2002.01340.x |
Gockel G, Hachtel W, Baier S, Fliss C, Henke M. 1994. Genes for components of the chloroplast translational apparatus are conserved in the reduced 73-kb plastid DNA of the nonphotosynthetic euglenoid flagellate Astasia longa. Curr. Genet, 26(3): 256-262.
DOI:10.1007/BF00309557 |
Gockel G, Hachtel W, Michael M. 2000. Complete gene map of the plastid genome of the nonphotosynthetic euglenoid flagellate Astasia longa. Protist, 151(4): 347-351.
DOI:10.1078/S1434-4610(04)70033-4 |
Hadariová L, Vesteg M, Birčák E, Schwartzbach S D, Krajčovič J. 2017. An intact plastid genome is essential for the survival of colorless Euglena longa but not Euglena gracilis. Curr. Genet, 63(2): 331-341.
DOI:10.1007/s00294-016-0641-z |
Hallick R B, Hong L, Drager R G, Favreau M R, Monfort A, Orsat B, Spielmann A, Stutz E. 1993. Complete sequence of Euglena gracilis chloroplast DNA. Nucleic Acids Res., 21(15): 3 537-3 544.
DOI:10.1093/nar/21.15.3537 |
Heizmann P, Doly J, Hussein Y, Nicolas P, Nigon V, Bernardi G. 1981. The chloroplast genome of bleached mutants of Euglena gracilis. Biochim. Biophys. Acta, 653(3): 412-415.
DOI:10.1016/0005-2787(81)90197-0 |
Heizmann P, Ravel-Chapuis P, Nigon V. 1982. Minicircular DNA having sequence homologies with chloroplast DNA in a bleached mutant of Euglena gracilis. Curr. Genet, 6(2): 119-122.
|
Heizmann P, Salvador G F, Nigon V. 1976. Occurrence of plastidial rRNAs and plastidial structures in bleached mutants of Euglena gracilis. Exp. Cell Res., 99(2): 253-260.
DOI:10.1016/0014-4827(76)90581-4 |
Hussein Y, Heizmann P, Nicolas P, Nigon V. 1982. Quantitative estimations of chloroplast DNA in bleached mutants of Euglena gracilis. Curr. Genet, 6(2): 111-117.
|
Idoine A D, Boulouis A, Rupprecht J, Bock R. 2014. The diurnal logic of the expression of the chloroplast genome in Chlamydomonas reinhardtii. PLoS One, 9(10): e108760.
DOI:10.1371/journal.pone.0108760 |
Kivic P A, Vesk M. 1972. Structure and function in the euglenoid eyespot apparatus: the fine structure, and response to environmental changes. Planta, 105(1): 1-14.
DOI:10.1007/BF00385158 |
Kivic P A, Vesk M. 1974. An electron microscope search for plastids in bleached Euglena gracilis and in Astasia longa. Can. J. Bot., 52(4): 695-699.
DOI:10.1139/b74-089 |
Krajčovič J, Ebringer L, Polónyi J. 1989. Quinolones and coumarins eliminate chloroplasts from Euglena gracilis. Antimicrob. Agents Chemother., 33(11): 1 883-1 889.
DOI:10.1128/AAC.33.11.1883 |
Krinsky N I, Goldsmith T H. 1960. The carotenoids of the flagellated alga, Euglena gracilis. Arch. Biochem. Biophys., 91(2): 271-279.
DOI:10.1016/0003-9861(60)90501-4 |
Križková L, Nagy M, Polónyi J, Ebringer L. 1998. The effect of flavonoids on ofloxacin-induced mutagenicity in Euglena gracilis. Mutat. Res., 416(1-2): 85-92.
DOI:10.1016/S1383-5718(98)00080-1 |
Krnáčová K, Rydlová I, Vinarčíková M, Krajčovič J, Vesteg M, Horváth A. 2015. Characterization of oxidative phosphorylation enzymes in Euglena gracilis and its white mutant strain WgmZOflL. FEBS Lett., 589(6): 687-694.
DOI:10.1016/j.febslet.2015.01.035 |
Oldenburg D J, Bendich A J. 2016. The linear plastid chromosomes of maize: terminal sequences, structures, and implications for DNA replication. Curr. Genet, 62(2): 431-442.
|
Qin H W, Chen L F, Lu N, Zhao Y H, Yuan X. 2012. Toxic effects of enrofloxacin on Scenedesmus obliquus. Front. Environ. Sci. Eng., 6(1): 107-116.
DOI:10.1007/s11783-011-0327-1 |
Schwartzbach S D, Schiff J A. 1974. Chloroplast and cytoplasmic ribosomes of Euglena: selective binding of dihydrostreptomycin to chloroplast ribosomes. J. Bacteriol., 120(1): 334-341.
DOI:10.1128/JB.120.1.334-341.1974 |
Siemeister G, Hachtel W. 1989. A circular 73 kb DNA from the colourless flagellate Astasia longa that resembles the chloroplast DNA of Euglena: restriction and gene map. Curr. Genet, 15(6): 435-441.
DOI:10.1007/BF00376801 |
Sulli C, Fang Z W, Muchhal U, Schwartzbach S D. 1999. Topology of Euglena chloroplast protein precursors within endoplasmic reticulum to Golgi to chloroplast transport vesicles. J. Biol. Chem., 274(1): 457-463.
DOI:10.1074/jbc.274.1.457 |
Thomas E J, Ortiz W. 1995. Loss of chloroplast transcripts for proteins associated with photosystem II: an early event during heat-bleaching in Euglena gracilis. Plant Mol. Biol., 27(2): 317-325.
DOI:10.1007/BF00020186 |
Tucci S, Vacula R, Krajčovič J, Proksch P, Martin W. 2010. Variability of wax ester fermentation in natural and bleached Euglena gracilis strains in response to oxygen and the elongase inhibitor flufenacet. J. Eukaryot. Microbiol., 57(1): 63-69.
DOI:10.1111/j.1550-7408.2009.00452.x |
Vacula R, Steiner J M, Krajčovič J, Ebringer L, Löffelhardt W. 2001. Plastid state- and light-dependent regulation of the expression of nucleus-encoded genes for chloroplast proteins in the flagellate Euglena gracilis. Folia Microbiol., 46(5): 433-441.
DOI:10.1007/BF02814435 |
Vesteg M, Vacula R, Burey S, Löffelhardt W, Drahovská H, Martin W, Krajčovič J. 2009. Expression of nucleus-encoded genes for chloroplast proteins in the flagellate Euglena gracilis. J. Eukaryot. Microbiol., 56(2): 159-166.
DOI:10.1111/j.1550-7408.2008.00383.x |
Wang J X, Shi Z X, Xu X D. 2002. Chloroplast-less mutants of two species of Euglena. Acta Hydrobiol. Sin., 26(2): 175-179.
|
Wang J X, Shi Z X, Xu X D. 2004. Residual plastids of bleached mutants of Euglena gracilis and their effects on the expression of nucleus-encoded genes. Prog. Nat. Sci., 14(3): 213-217.
DOI:10.1080/10020070412331343371 |
Yoshida Y, Tomiyama T, Maruta T, Tomita M, Ishikawa T, Arakawa K. 2016. De novo assembly and comparative transcriptome analysis of Euglena gracilis in response to anaerobic conditions. BMC Genomics, 17: 182.
DOI:10.1186/s12864-016-2540-6 |
Zakryś B, Milanowski R, Karnkowska A. 2017. Evolutionary origin of Euglena. In: Schwartzbach S D, Shigeoka S eds. Euglena: Biochemistry, Cell and Molecular Biology. Springer, Cham. p.3-17.
|
 2020, Vol. 38
2020, Vol. 38